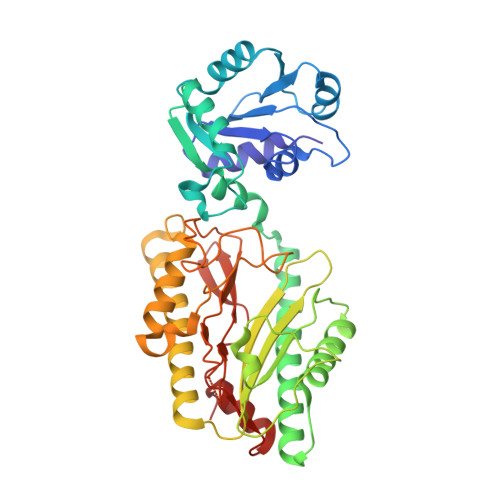
Crystal structure of a novel prolidase from Deinococcus radiodurans identifies new subfamily of bacterial prolidases.
Are, V.N., Jamdar, S.N., Ghosh, B., Goyal, V.D., Kumar, A., Neema, S., Gadre, R., Makde, R.D.(2017) Proteins 85: 2239-2251
- PubMed: 28929533
- DOI: https://doi.org/10.1002/prot.25389
- Primary Citation of Related Structures:
5XEV - PubMed Abstract:
Xaa-Pro peptidases (XPP) are dinuclear peptidases of MEROPS M24B family that hydrolyze Xaa-Pro iminopeptide bond with a trans-proline at the second position of the peptide substrate. XPPs specific towards dipeptides are called prolidases while those that prefer longer oligopeptides are called aminopeptidases P. Though XPPs are strictly conserved in bacterial and archaeal species, the structural and sequence features that distinguish between prolidases and aminopeptidases P are not always clear. Here, we report 1.4 Å resolution crystal structure of a novel XPP from Deinococcus radiodurans (XPPdr). XPPdr forms a novel dimeric structure via unique dimer stabilization loops of N-terminal domains such that their C-terminal domains are placed far apart from each other. This novel dimerization is also the consequence of a different orientation of N-terminal domain in XPPdr monomer than those in other known prolidases. The enzymatic assays show that it is a prolidase with broad substrate specificity. Our structural, mutational, and molecular dynamics simulation analyses show that the conserved Arg46 of N-terminal domain is important for the dipeptide selectivity. Our BLAST search found XPPdr orthologs with conserved sequence motifs which correspond to unique structural features of XPPdr, thus identify a new subfamily of bacterial prolidases.
Organizational Affiliation:
High Pressure and Synchrotron Radiation Physics Division, Bhabha Atomic Research Centre, Mumbai, India.