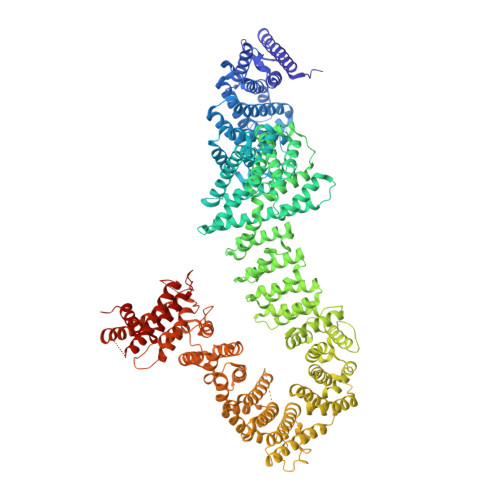
Crystal structure of the cohesin loader Scc2 and insight into cohesinopathy.
Kikuchi, S., Borek, D.M., Otwinowski, Z., Tomchick, D.R., Yu, H.(2016) Proc Natl Acad Sci U S A 113: 12444-12449
- PubMed: 27791135
- DOI: https://doi.org/10.1073/pnas.1611333113
- Primary Citation of Related Structures:
5T8V - PubMed Abstract:
The ring-shaped cohesin complex topologically entraps chromosomes and regulates chromosome segregation, transcription, and DNA repair. The cohesin core consists of the structural maintenance of chromosomes 1 and 3 (Smc1-Smc3) heterodimeric ATPase, the kleisin subunit sister chromatid cohesion 1 (Scc1) that links the two ATPase heads, and the Scc1-bound adaptor protein Scc3. The sister chromatid cohesion 2 and 4 (Scc2-Scc4) complex loads cohesin onto chromosomes. Mutations of cohesin and its regulators, including Scc2, cause human developmental diseases termed cohesinopathy. Here, we report the crystal structure of Chaetomium thermophilum (Ct) Scc2 and examine its interaction with cohesin. Similar to Scc3 and another Scc1-interacting cohesin regulator, precocious dissociation of sisters 5 (Pds5), Scc2 consists mostly of helical repeats that fold into a hook-shaped structure. Scc2 binds to Scc1 through an N-terminal region of Scc1 that overlaps with its Pds5-binding region. Many cohesinopathy mutations target conserved residues in Scc2 and diminish Ct Scc2 binding to Ct Scc1. Pds5 binding to Scc1 weakens the Scc2-Scc1 interaction. Our study defines a functionally important interaction between the kleisin subunit of cohesin and the hook of Scc2. Through competing with Scc2 for Scc1 binding, Pds5 might contribute to the release of Scc2 from loaded cohesin, freeing Scc2 for additional rounds of loading.
Organizational Affiliation:
Department of Pharmacology, University of Texas Southwestern Medical Center, Dallas, TX 75390.