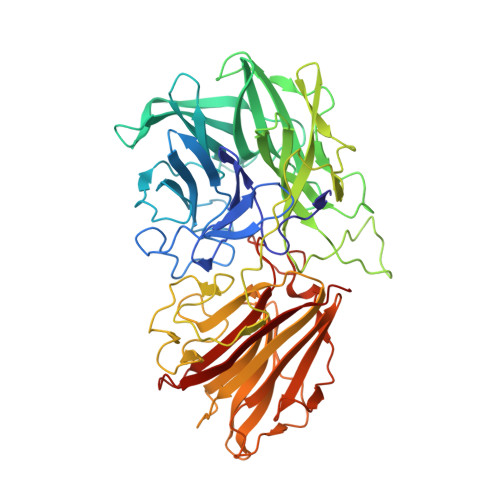
Structure-based protein engineering of bacterial beta-xylosidase to increase the production yield of xylobiose from xylose
Hong, S., Kyung, M., Jo, I., Kim, Y.R., Ha, N.C.(2018) Biochem Biophys Res Commun 501: 703-710
- PubMed: 29752942
- DOI: https://doi.org/10.1016/j.bbrc.2018.05.051
- Primary Citation of Related Structures:
5ZQJ, 5ZQS, 5ZQX - PubMed Abstract:
Xylobiose consists of two molecules of xylose and has been highly recognized as a food supplement because it possesses high prebiotic functions. β-xylosidase exhibits enzymatic activity to hydrolyze xylobiose, and the enzyme can also catalyze the reverse reaction in the presence of high concentrations of xylose. Previously, β-xylosidase from Bacillus pumilus IPO (BpXynB), belonging to GH family 43, was employed to produce xylobiose from xylose. To improve the enzymatic efficiency, this study determined the high-resolution structure of BpXynB in a complex with xylobiose and engineered BpXynB based on the structures. The structure of BpXynB deciphered the residues involved in the recognition of the xylobiose. A site-directed mutation at the residue for xylobiose recognition increased the yield of xylobiose by 20% compared to a similar activity of the wild type enzyme. The complex structure of the mutant enzyme and xylobiose provided the structural basis for a higher yield of the engineered protein. This engineered enzyme would enable a higher economic production of xylobiose, and a similar engineering strategy could be applied within the same family of enzymes.
Organizational Affiliation:
Research Institute for Agriculture and Life Sciences, Center for Food and Bioconvergence, Center for Food Safety and Toxicology, Department of Agicultural Biotechnology, Seoul National University, Seoul, 08826, Republic of Korea.