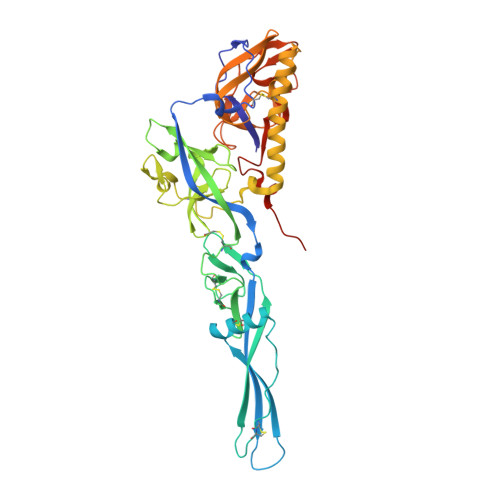
Structural intermediates in the fusion-associated transition of vesiculovirus glycoprotein.
Baquero, E., Albertini, A.A., Raux, H., Abou-Hamdan, A., Boeri-Erba, E., Ouldali, M., Buonocore, L., Rose, J.K., Lepault, J., Bressanelli, S., Gaudin, Y.(2017) EMBO J 36: 679-692
- PubMed: 28188244
- DOI: https://doi.org/10.15252/embj.201694565
- Primary Citation of Related Structures:
5MDM - PubMed Abstract:
Vesiculoviruses enter cells by membrane fusion, driven by a large, low-pH-induced, conformational change in the fusion glycoprotein G that involves transition from a trimeric pre-fusion toward a trimeric post-fusion state via monomeric intermediates. Here, we present the structure of the G fusion protein at intermediate pH for two vesiculoviruses, vesicular stomatitis virus (VSV) and Chandipura virus (CHAV), which is responsible for deadly encephalopathies. First, a CHAV G crystal structure shows two intermediate conformations forming a flat dimer of heterodimers. On virions, electron microscopy (EM) and tomography reveal monomeric spikes similar to one of the crystal conformations. In solution, mass spectrometry shows dimers of G. Finally, mutations at a dimer interface, involving fusion domains associated in an antiparallel manner to form an intermolecular β-sheet, affect G fusion properties. The location of the compensatory mutations restoring fusion activity strongly suggests that this interface is functionally relevant. This work reveals the range of G structural changes and suggests that G monomers can re-associate, through antiparallel interactions between fusion domains, into dimers that play a role at some early stage of the fusion process.
Organizational Affiliation:
Institute for Integrative Biology of the Cell (I2BC), CEA, CNRS, Univ. Paris-Sud, Université Paris-Saclay, Gif-sur-Yvette Cedex, France.