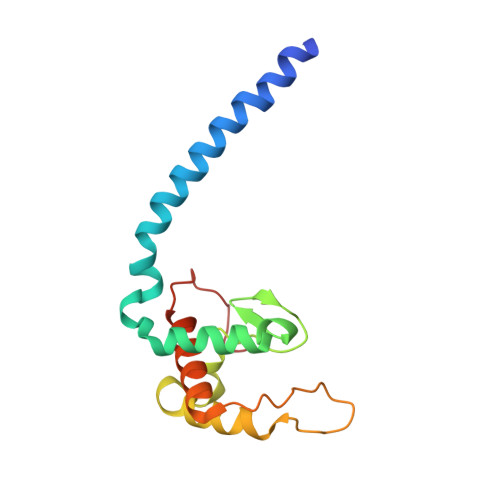
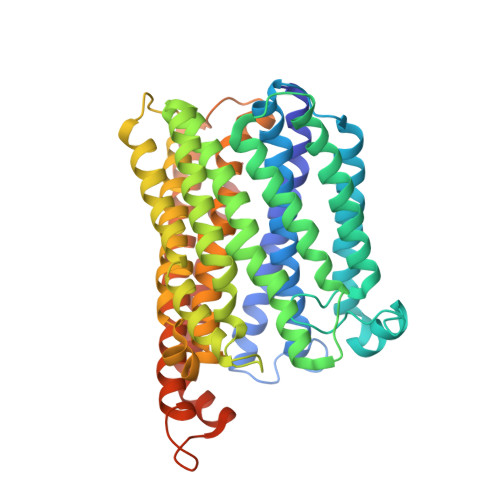
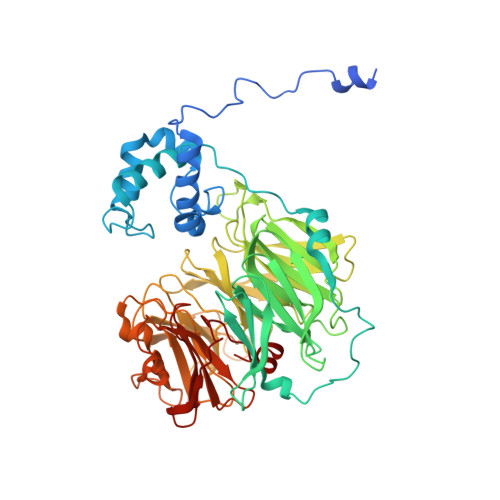
Dynamics of nitric oxide controlled by protein complex in bacterial system
Terasaka, E., Yamada, K., Wang, P.H., Hosokawa, K., Yamagiwa, R., Matsumoto, K., Ishii, S., Mori, T., Yagi, K., Sawai, H., Arai, H., Sugimoto, H., Sugita, Y., Shiro, Y., Tosha, T.(2017) Proc Natl Acad Sci U S A 114: 9888-9893
- PubMed: 28847930
- DOI: https://doi.org/10.1073/pnas.1621301114
- Primary Citation of Related Structures:
5GUW, 5GUX - PubMed Abstract:
Nitric oxide (NO) plays diverse and significant roles in biological processes despite its cytotoxicity, raising the question of how biological systems control the action of NO to minimize its cytotoxicity in cells. As a great example of such a system, we found a possibility that NO-generating nitrite reductase (NiR) forms a complex with NO-decomposing membrane-integrated NO reductase (NOR) to efficiently capture NO immediately after its production by NiR in anaerobic nitrate respiration called denitrification. The 3.2-Å resolution structure of the complex of one NiR functional homodimer and two NOR molecules provides an idea of how these enzymes interact in cells, while the structure may not reflect the one in cells due to the membrane topology. Subsequent all-atom molecular dynamics (MD) simulations of the enzyme complex model in a membrane and structure-guided mutagenesis suggested that a few interenzyme salt bridges and coulombic interactions of NiR with the membrane could stabilize the complex of one NiR homodimer and one NOR molecule and contribute to rapid NO decomposition in cells. The MD trajectories of the NO diffusion in the NiR:NOR complex with the membrane showed that, as a plausible NO transfer mechanism, NO released from NiR rapidly migrates into the membrane, then binds to NOR. These results help us understand the mechanism of the cellular control of the action of cytotoxic NO.
Organizational Affiliation:
Biometal Science Laboratory, RIKEN SPring-8 Center, Kouto, Sayo, Hyogo 679-5148, Japan.