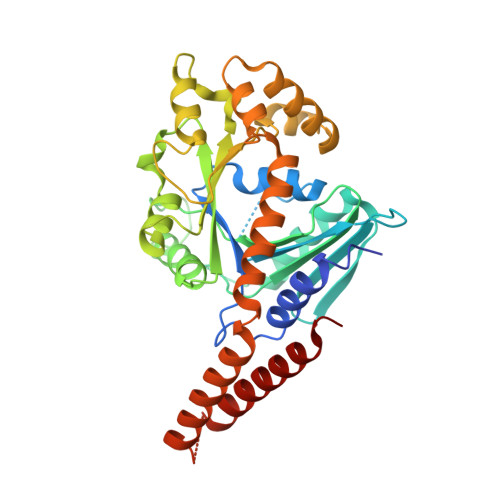
Transient Dimerization of Human MxA Promotes GTP Hydrolysis, Resulting in a Mechanical Power Stroke.
Rennie, M.L., McKelvie, S.A., Bulloch, E.M., Kingston, R.L.(2014) Structure 22: 1433-1445
- PubMed: 25295396
- DOI: https://doi.org/10.1016/j.str.2014.08.015
- Primary Citation of Related Structures:
4P4S, 4P4T, 4P4U - PubMed Abstract:
Myxovirus resistance (Mx) proteins restrict replication of numerous viruses. They are closely related to membrane-remodeling fission GTPases, such as dynamin. Mx proteins can tubulate lipids and form rings or filaments that may interact directly with viral structures. GTPase domain dimerization is thought to allow crosstalk between the rungs of a tubular or helical assembly, facilitating constriction. We demonstrate that the GTPase domain of MxA dimerizes to facilitate catalysis, in a fashion analogous to dynamin. GTP binding is associated with the lever-like movement of structures adjacent to the GTPase domain, while GTP hydrolysis returns MxA to its resting state. Dimerization is not significantly promoted by substrate binding and occurs only transiently, yet is central to catalytic efficiency. Therefore, we suggest dimerization functions to coordinate the activity of spatially adjacent Mx molecules within an assembly, allowing their mechanical power strokes to be synchronized at key points in the contractile cycle.
Organizational Affiliation:
Maurice Wilkins Centre, School of Biological Sciences, University of Auckland, Auckland 1010, New Zealand.