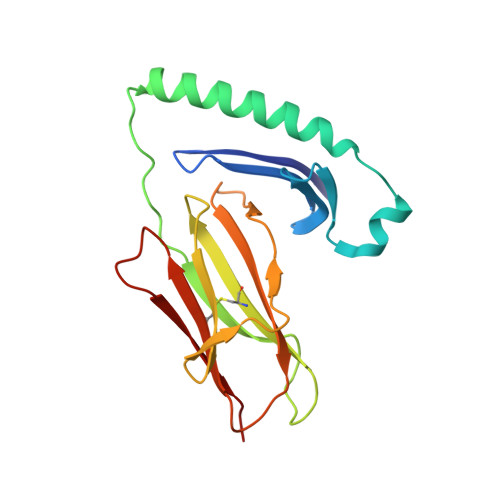
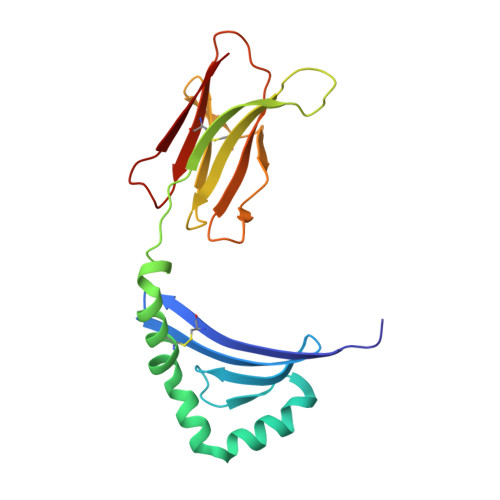
T-cell receptor (TCR) interaction with peptides that mimic nickel offers insight into nickel contact allergy.
Yin, L., Crawford, F., Marrack, P., Kappler, J.W., Dai, S.(2012) Proc Natl Acad Sci U S A 109: 18517-18522
- PubMed: 23091041
- DOI: https://doi.org/10.1073/pnas.1215928109
- Primary Citation of Related Structures:
4H1L, 4H25, 4H26 - PubMed Abstract:
T cell-mediated allergy to Ni(++) is one of the most common forms of allergic contact dermatitis, but how the T-cell receptor (TCR) recognizes Ni(++) is unknown. We studied a TCR from an allergic patient that recognizes Ni(++) bound to the MHCII molecule DR52c containing an unknown self-peptide. We identified mimotope peptides that can replace both the self-peptide and Ni(++) in this ligand. They share a p7 lysine whose εNH(2) group is surface-exposed when bound to DR52c. Whereas the TCR uses germ-line complementary-determining region (CDR)1/2 amino acids to dock in the conventional diagonal mode on the mimotope-DR52c complex, the interface is dominated by the TCR Vβ CDR3 interaction with the p7 lysine. Mutations in the TCR CDR loops have similar effects on the T-cell response to either the mimotope or Ni(++) ligand. We suggest that the mimotope p7 lysine mimics Ni(++) in the natural TCR ligand and that MHCII β-chain flexibility in the area around the peptide p7 position forms a common site for cation binding in metal allergies.
Organizational Affiliation:
Howard Hughes Medical Institute, National Jewish Health, Denver, CO 80206, USA.