Structures of Cd200/Cd200 Receptor Family and Implications for Topology, Regulation, and Evolution
Hatherley, D., Lea, S.M., Johnson, S., Barclay, A.N.(2013) Structure 21: 820
- PubMed: 23602662
- DOI: https://doi.org/10.1016/j.str.2013.03.008
- Primary Citation of Related Structures:
4BFE, 4BFG, 4BFI - PubMed Abstract:
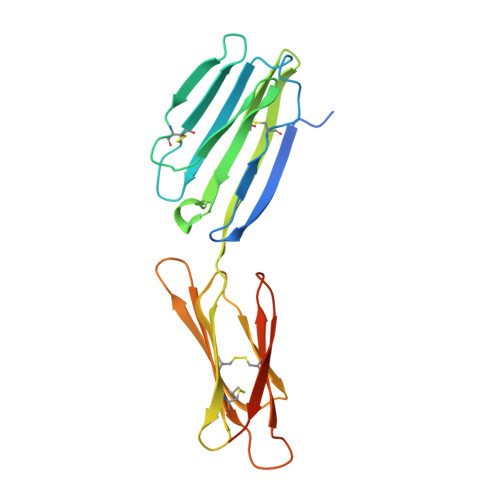
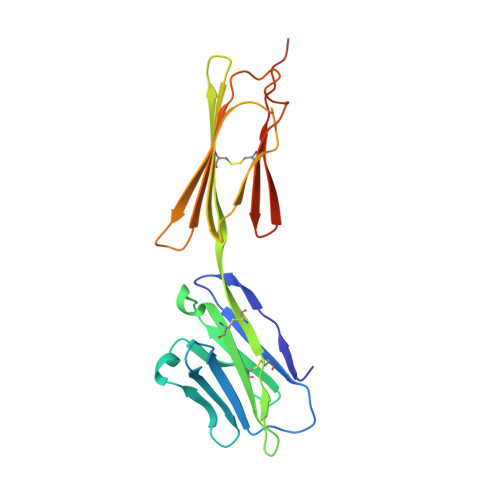
CD200 is a widely distributed membrane glycoprotein that regulates myeloid cell activity through its interaction with an inhibitory receptor (CD200R). The interaction is of interest as a target for treating excessive inflammation and for treating leukemia. There are closely related proteins to CD200R that give activating signals making this a "paired receptor." We report X-ray crystallography structures for the inhibitory CD200R, the activating receptor CD200RLa, and a complex between CD200R and CD200. Both CD200 and CD200R contain two Ig-like domains and interact through their NH₂ terminal domains compatible with immunological synapse-like interactions occurring between myeloid cells and other CD200-expressing cells. The failure of the activating receptor to bind CD200 resides in subtle changes around the interface. CD200 has been acquired by herpes viruses to mimic the host interaction. CD200R has evolved rapidly presumably driven by pathogen pressure but it may also be important in homeostasis through interactions with commensal bacteria.
Organizational Affiliation:
Sir William Dunn School of Pathology, University of Oxford, South Parks Road, Oxford OX1 3RE, UK.