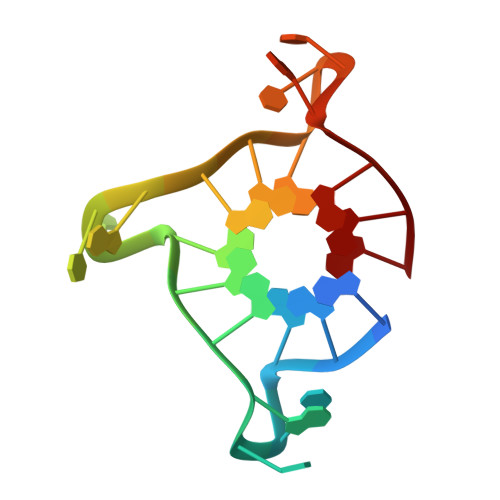
Structure-based design and evaluation of naphthalene diimide g-quadruplex ligands as telomere targeting agents in pancreatic cancer cells.
Micco, M., Collie, G.W., Dale, A.G., Ohnmacht, S.A., Pazitna, I., Gunaratnam, M., Reszka, A.P., Neidle, S.(2013) J Med Chem 56: 2959-2974
- PubMed: 23514618
- DOI: https://doi.org/10.1021/jm301899y
- Primary Citation of Related Structures:
3UYH, 4DA3, 4DAQ - PubMed Abstract:
Tetra-substituted naphthalene diimide (ND) derivatives with positively charged termini are potent stabilizers of human telomeric and gene promoter DNA quadruplexes and inhibit the growth of human cancer cells in vitro and in vivo. The present study reports the enhancement of the pharmacological properties of earlier ND compounds using structure-based design. Crystal structures of three complexes with human telomeric intramolecular quadruplexes demonstrate that two of the four strongly basic N-methyl-piperazine groups can be replaced by less basic morpholine groups with no loss of intermolecular interactions in the grooves of the quadruplex. The new compounds retain high affinity to human telomeric quadruplex DNA but are 10-fold more potent against the MIA PaCa-2 pancreatic cancer cell line, with IC50 values of ~10 nM. The lead compound induces cellular senescence but does not inhibit telomerase activity at the nanomolar dosage levels required for inhibition of cellular proliferation. Gene array qPCR analysis of MIA PaCa-2 cells treated with the lead compound revealed significant dose-dependent modulation of a distinct subset of genes, including strong induction of DNA damage responsive genes CDKN1A, DDIT3, GADD45A/G, and PPM1D, and repression of genes involved in telomere maintenance, including hPOT1 and PARP1.
Organizational Affiliation:
The School of Pharmacy, University College London, London WC1N 1AX, UK.