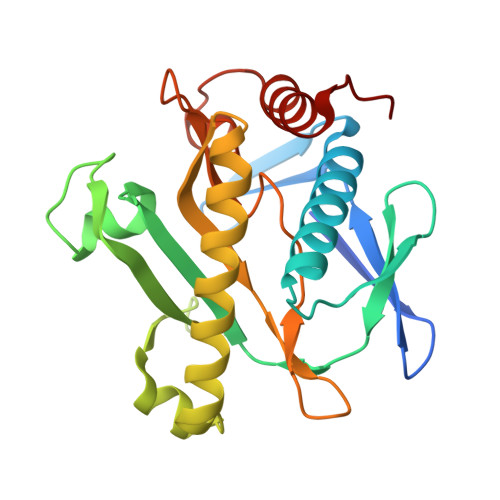
Structure of SAICAR synthetase from Pyrococcus horikoshii OT3: insights into thermal stability
Manjunath, K., Kanaujia, S.P., Kanagaraj, S., Jeyakanthan, J., Sekar, K.(2013) Int J Biol Macromol 53: 7-19
- PubMed: 23137517
- DOI: https://doi.org/10.1016/j.ijbiomac.2012.10.028
- Primary Citation of Related Structures:
3U54, 3U55 - PubMed Abstract:
The first native crystal structure of Phosphoribosylaminoimidazole-succinocarboxamide synthetase (SAICAR synthetase) from a hyperthermophilic organism Pyrococcus horikoshii OT3 was determined in two space groups H3 (Type-1: Resolution 2.35Å) and in C222(1) (Type-2: Resolution 1.9Å). Both are dimeric but Type-1 structure exhibited hexameric arrangement due to the presence of cadmium ions. A comparison has been made on the sequence and structures of all SAICAR synthetases to better understand the differences between mesophilic, thermophilic and hyperthermophilic SAICAR synthetases. These SAICAR synthetases are reasonably similar in sequence and three-dimensional structure; however, differences were visible only in the subtler details of percentage composition of the sequences, salt bridge interactions and non-polar contact areas.
Organizational Affiliation:
Supercomputer Education and Research Centre, Indian Institute of Science, Bangalore 560 012, India.