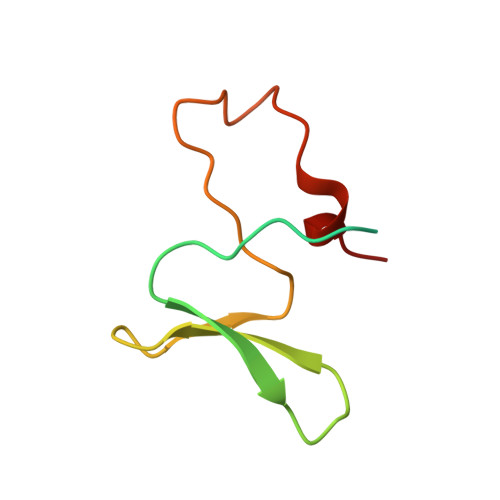
Structure of the dimerization domain of DiGeorge Critical Region 8
Senturia, R., Faller, M., Yin, S., Loo, J.A., Cascio, D., Sawaya, M.R., Hwang, D., Clubb, R.T., Guo, F.(2010) Protein Sci 19: 1354-1365
- PubMed: 20506313
- DOI: https://doi.org/10.1002/pro.414
- Primary Citation of Related Structures:
3LE4 - PubMed Abstract:
Maturation of microRNAs (miRNAs, approximately 22nt) from long primary transcripts [primary miRNAs (pri-miRNAs)] is regulated during development and is altered in diseases such as cancer. The first processing step is a cleavage mediated by the Microprocessor complex containing the Drosha nuclease and the RNA-binding protein DiGeorge critical region 8 (DGCR8). We previously reported that dimeric DGCR8 binds heme and that the heme-bound DGCR8 is more active than the heme-free form. Here, we identified a conserved dimerization domain in DGCR8. Our crystal structure of this domain (residues 298-352) at 1.7 A resolution demonstrates a previously unknown use of a WW motif as a platform for extensive dimerization interactions. The dimerization domain of DGCR8 is embedded in an independently folded heme-binding domain and directly contributes to association with heme. Heme-binding-deficient DGCR8 mutants have reduced pri-miRNA processing activity in vitro. Our study provides structural and biochemical bases for understanding how dimerization and heme binding of DGCR8 may contribute to regulation of miRNA biogenesis.
Organizational Affiliation:
Department of Biological Chemistry in David Geffen School of Medicine, University of California, Los Angeles, California 90095, USA.