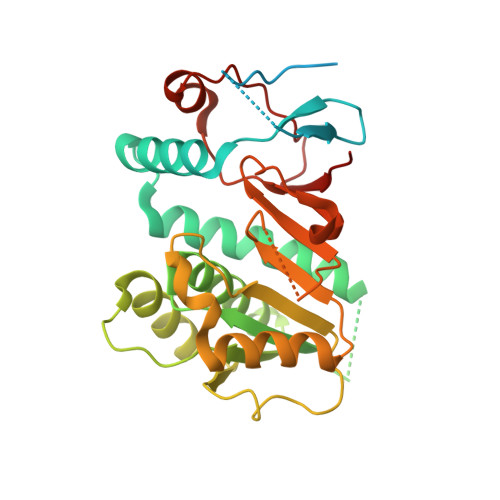
Crystal structure of human Edc3 and its functional implications
Ling, S.H.M., Decker, C.J., Walsh, M.A., She, M., Parker, R., Song, H.(2008) Mol Cell Biol 28: 5965-5976
- PubMed: 18678652
- DOI: https://doi.org/10.1128/MCB.00761-08
- Primary Citation of Related Structures:
3D3J, 3D3K - PubMed Abstract:
Edc3 is an enhancer of decapping and serves as a scaffold that aggregates mRNA ribonucleoproteins together for P-body formation. Edc3 forms a network of interactions with the components of the mRNA decapping machinery and has a modular domain architecture consisting of an N-terminal Lsm domain, a central FDF domain, and a C-terminal YjeF-N domain. We have determined the crystal structure of the N-terminally truncated human Edc3 at a resolution of 2.2 A. The structure reveals that the YjeF-N domain of Edc3 possesses a divergent Rossmann fold topology that forms a dimer, which is supported by sedimentation velocity and sedimentation equilibrium analysis in solution. The dimerization interface of Edc3 is highly conserved in eukaryotes despite the overall low sequence homology across species. Structure-based site-directed mutagenesis revealed dimerization is required for efficient RNA binding, P-body formation, and likely for regulating the yeast Rps28B mRNA as well, suggesting that the dimeric form of Edc3 is a structural and functional unit in mRNA degradation.
Organizational Affiliation:
Laboratory of Macromolecular Structure, Institute of Molecular and Cell Biology, 61 Biopolis Drive, Singapore 138673, Singapore.