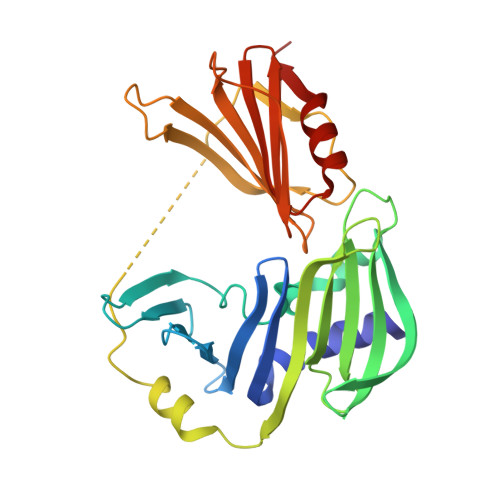
Crystal structure of RseB and a model of its binding mode to RseA
Kim, D.Y., Jin, K.S., Kwon, E., Ree, M., Kim, K.K.(2007) Proc Natl Acad Sci U S A 104: 8779-8784
- PubMed: 17496148
- DOI: https://doi.org/10.1073/pnas.0703117104
- Primary Citation of Related Structures:
2P4B - PubMed Abstract:
The bacterial envelope stress response senses stress signals in the extracytoplasmic compartment, and activates sigma(E)-dependent transcription by degrading its antisigma factor RseA. RseB, a binding partner of RseA, plays a pivotal role in regulating this response, but its molecular mechanism is not understood. We therefore determined the crystal structure of Escherichia coli RseB at a resolution of 2.4 A. RseB is composed of two domains linked by a flexible linker and forms a loosely packed dimer with two grooves on each side. This structural feature is confirmed by small-angle scattering in solution. Analysis of the binding of various RseA mutants to RseB allowed us to identify the major RseB-binding motif in RseA. These data, coupled with analysis of small-angle scattering of the RseA/RseB complex in solution, leads us to propose that two RseAs bind to the grooves of the dimeric RseB by conserved residues. The implications for modulating proteolytic cleavage of RseA are discussed.
Organizational Affiliation:
Department of Molecular Cell Biology, Samsung Biomedical Research Institute, Sungkyunkwan University School of Medicine, Suwon 440-746, Korea.