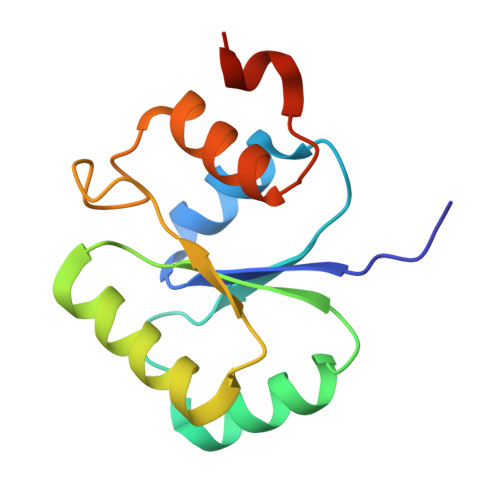
Crystal structure and putative function of small Toprim domain-containing protein from Bacillus stearothermophilus.
Rezacova, P., Borek, D., Moy, S.F., Joachimiak, A., Otwinowski, Z.(2008) Proteins 70: 311-319
- PubMed: 17705269
- DOI: https://doi.org/10.1002/prot.21511
- Primary Citation of Related Structures:
2FCJ, 2I5R - PubMed Abstract:
The crystal structure of the Midwest Center for Structural Genomics target APC35832, a 14.7-kDa cytosolic protein from Bacillus stearothermophilus, has been determined at 1.3 A resolution by the single anomalous diffraction method from a mercury soaked crystal. The APC35832 protein is a representative of large group of bacterial and archeal proteins entirely consisting of the Toprim (topoisomerase-primase) domain. This domain is found in the catalytic centers of many enzymes catalyzing phosphodiester bond formation or cleavage, but the function of small Toprim domain proteins remains unknown. Consistent with the sequence analysis, the APC35832 structure shows a conserved Toprim fold, with a central 4-stranded parallel beta-sheet surrounded by four alpha-helixes. Comparison of the APC35832 structure with its closest structural homolog, the catalytic core of bacteriophage T7 primase, revealed structural conservation of a metal binding site and isothermal titration calorimetry indicates that APC35832 binds Mg2+ with a sub-millimolar dissociation constant (K(d)). The APC35832-Mg2+ complex structure was determined at 1.65 A and reveals the role of conserved acidic residues in Mg2+ ion coordination. The structural similarities to other Toprim domain containing proteins and potential function and substrates of APC35832 are discussed in this article.
Organizational Affiliation:
Department of Biochemistry, UT Southwestern Medical Center, Dallas, Texas 75390-8816, USA.