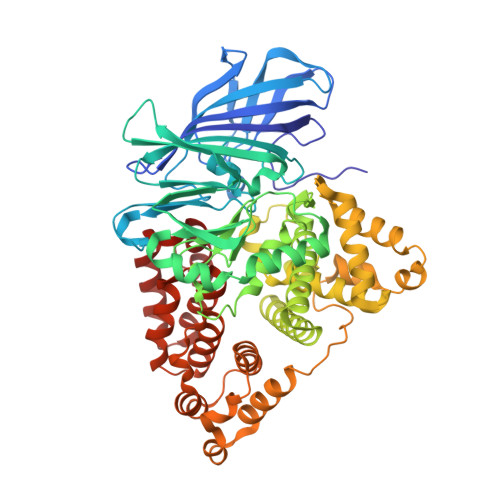
Leukotriene A4 hydrolase: identification of a common carboxylate recognition site for the epoxide hydrolase and aminopeptidase substrates
Rudberg, P.C., Tholander, F.O.T., Andberg, M., Thunnissen, M.M.G.M.(2004) J Biol Chem 279: 27376-27382
- PubMed: 15078870
- DOI: https://doi.org/10.1074/jbc.M401031200
- Primary Citation of Related Structures:
1SQM - PubMed Abstract:
Leukotriene (LT) A(4) hydrolase is a bifunctional zinc metalloenzyme, which converts LTA(4) into the neutrophil chemoattractant LTB(4) and also exhibits an anion-dependent aminopeptidase activity. In the x-ray crystal structure of LTA(4) hydrolase, Arg(563) and Lys(565) are found at the entrance of the active center. Here we report that replacement of Arg(563), but not Lys(565), leads to complete abrogation of the epoxide hydrolase activity. However, mutations of Arg(563) do not seem to affect substrate binding strength, because values of K(i) for LTA(4) are almost identical for wild type and (R563K)LTA(4) hydrolase. These results are supported by the 2.3-A crystal structure of (R563A)LTA(4) hydrolase, which does not reveal structural changes that can explain the complete loss of enzyme function. For the aminopeptidase reaction, mutations of Arg(563) reduce the catalytic activity (V(max) = 0.3-20%), whereas mutations of Lys(565) have limited effect on catalysis (V(max) = 58-108%). However, in (K565A)- and (K565M)LTA(4) hydrolase, i.e. mutants lacking a positive charge, values of the Michaelis constant for alanine-p-nitroanilide increase significantly (K(m) = 480-640%). Together, our data indicate that Arg(563) plays an unexpected, critical role in the epoxide hydrolase reaction, presumably in the positioning of the carboxylate tail to ensure perfect substrate alignment along the catalytic elements of the active site. In the aminopeptidase reaction, Arg(563) and Lys(565) seem to cooperate to provide sufficient binding strength and productive alignment of the substrate. In conclusion, Arg(563) and Lys(565) possess distinct roles as carboxylate recognition sites for two chemically different substrates, each of which is turned over in separate enzymatic reactions catalyzed by LTA(4) hydrolase.
Organizational Affiliation:
Department of Medical Biochemistry and Biophysics, Division of Chemistry II, Karolinska Institutet, S-171 77 Stockholm, Sweden.