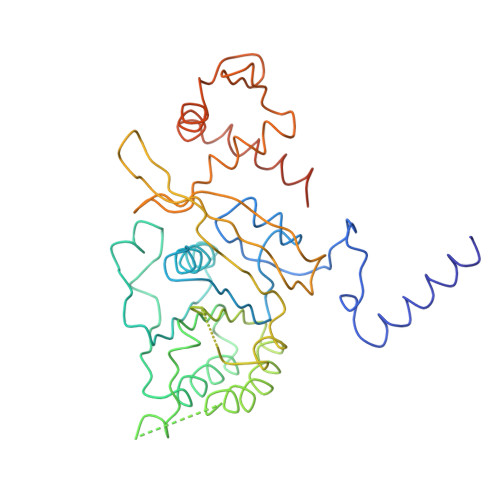
Structure of the recA protein-ADP complex.
Story, R.M., Steitz, T.A.(1992) Nature 355: 374-376
- PubMed: 1731253
- DOI: https://doi.org/10.1038/355374a0
- Primary Citation of Related Structures:
1REA - PubMed Abstract:
The recA protein catalyses the ATP-driven homologous pairing and strand exchange of DNA molecules. It is an allosteric enzyme: the ATPase activity is DNA-dependent, and ATP-bound recA protein has a high affinity for DNA, whereas the ADP-bound form has a low affinity. In the absence of ATP hydrolysis, recA protein can still promote homologous pairing, apparently through the formation of a triple-stranded intermediate. The exact role of ATP hydrolysis is not clear, but it presumably drives the triplex intermediate towards products. Here we determine the position of bound ADP diffused into the recA crystal. We show that only the phosphates are bound in the same way as in other NTPases containing the G/AXXXXGKT/S motif. We propose that recA protein may change its conformation upon ATP hydrolysis in a manner analogous to one such protein, the p21 protein from the ras oncogene. A model is presented to account for the allosteric stimulation of DNA binding by ATP. The mechanism by which nucleoside triphosphate hydrolysis is coupled to the binding of another ligand in recA protein and p21 may be typical of the large class of NTPases containing this conserved motif.
Organizational Affiliation:
Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, Connecticut 06511.