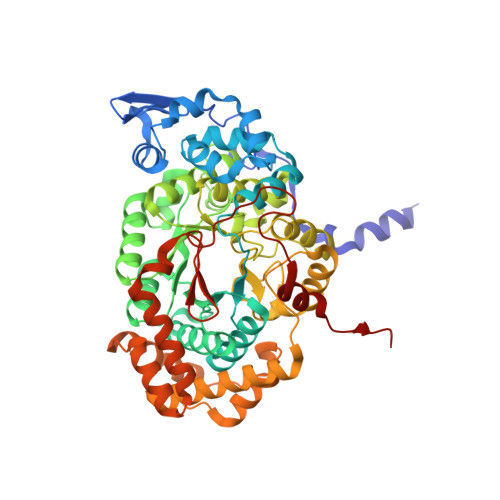
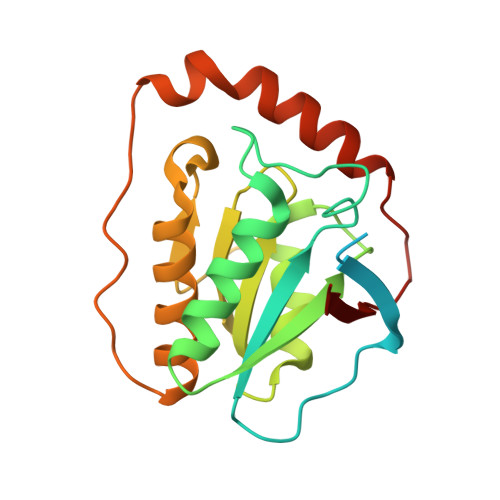
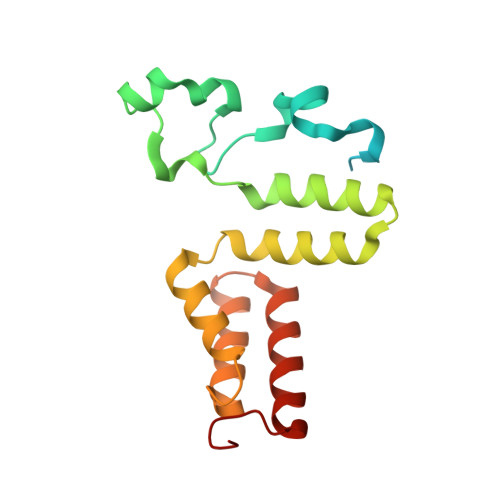
Substrate-induced conformational change of a coenzyme B12-dependent enzyme: crystal structure of the substrate-free form of diol dehydratase
Shibata, N., Masuda, J., Morimoto, Y., Yasuoka, N., Toraya, T.(2002) Biochemistry 41: 12607-12617
- PubMed: 12379103
- DOI: https://doi.org/10.1021/bi026104z
- Primary Citation of Related Structures:
1IWB - PubMed Abstract:
Substrate binding triggers catalytic radical formation through the cobalt-carbon bond homolysis in coenzyme B12-dependent enzymes. We have determined the crystal structure of the substrate-free form of Klebsiella oxytoca diol dehydratase*cyanocobalamin complex at 1.85 A resolution. The structure contains two units of the heterotrimer consisting of alpha, beta, and gamma subunits. As compared with the structure of its substrate-bound form, the beta subunits are tilted by approximately 3 degrees and cobalamin is also tilted so that pyrrole rings A and D are significantly lifted up toward the substrate-binding site, whereas pyrrole rings B and C are only slightly lifted up. The structure revealed that the potassium ion in the substrate-binding site of the substrate-free enzyme is also heptacoordinated; that is, two oxygen atoms of two water molecules coordinate to it instead of the substrate hydroxyls. A modeling study in which the structures of both the cobalamin moiety and the adenine ring of the coenzyme were superimposed onto those of the enzyme-bound cyanocobalamin and the adenine ring-binding pocket, respectively, demonstrated that the distortions of the Co-C bond in the substrate-free form are already marked but slightly smaller than those in the substrate-bound form. It was thus strongly suggested that the Co-C bond becomes largely activated (labilized) when the coenzyme binds to the apoenzyme even in the absence of substrate and undergoes homolysis through the substrate-induced conformational changes of the enzyme. Kinetic coupling of Co-C bond homolysis with hydrogen abstraction from the substrate shifts the equilibrium to dissociation.
Organizational Affiliation:
Department of Life Science, Graduate School of Science, Himeji Institute of Technology, 3-2-1 Kouto, Kamigori, Ako-gun, Hyogo 678-1297, Japan.