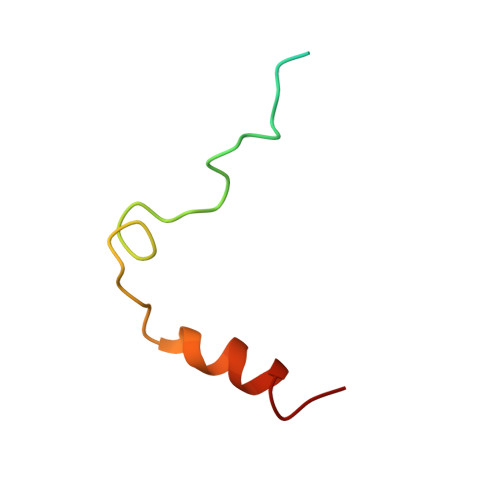
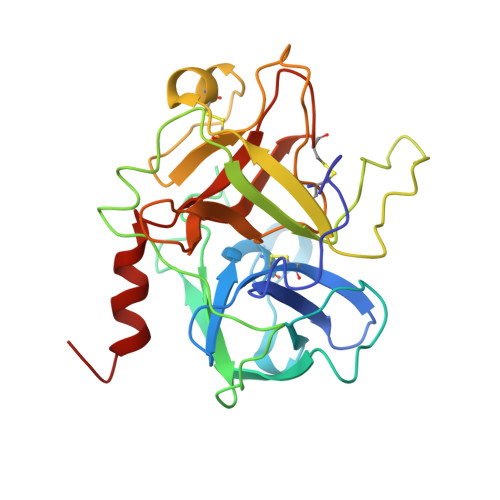
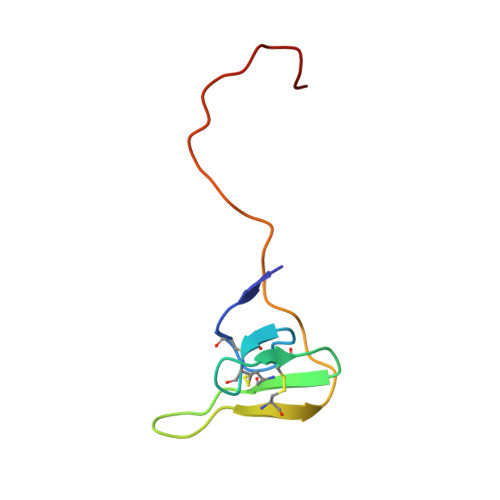
The structure of a complex of bovine alpha-thrombin and recombinant hirudin at 2.8-A resolution.
Vitali, J., Martin, P.D., Malkowski, M.G., Robertson, W.D., Lazar, J.B., Winant, R.C., Johnson, P.H., Edwards, B.F.(1992) J Biol Chem 267: 17670-17678
- PubMed: 1517214
- Primary Citation of Related Structures:
1HRT - PubMed Abstract:
Crystals of the complex of bovine alpha-thrombin with recombinant hirudin variant 1 have space group C222(1) with cell constants a = 59.11, b = 102.62, and c = 143.26 A. The orientation and position of the thrombin component was determined by molecular replacement and the hirudin molecule was fit in 2 magnitude of Fo - magnitude of Fc electron density maps. The structure was refined by restrained least squares and simulated annealing to R = 0.161 at 2.8-A resolution. The binding of hirudin to thrombin is generally similar to that observed in the crystals of human thrombin-hirudin. Several differences in the interactions of the COOH-terminal polypeptide of hirudin, specifically of residues Asp-55h, Phe-56h, Glu-57h, and Glu-58h, and a few differences in the interactions of the hirudin core, specifically of residues Asp-5h, Ser-19h, and Asn-20h, with thrombin from human thrombin-hirudin suggest that there is some flexibility in the binding of these 2 molecules. Most of the residues in the 9 subsites that bind fibrinopeptide A7-16 to thrombin also interact with the NH2-terminal domain of hirudin. The S1 subsite is a notable exception in that only 1 of its 6 residues, namely Ser-214, interacts with hirudin. The only difference between human and bovine thrombins that appears to influence the binding of hirudin is the replacement of Lys-149E by an acidic glutamate in the bovine enzyme.
Organizational Affiliation:
Department of Biochemistry, Wayne State University, Detroit, Michigan 48201.