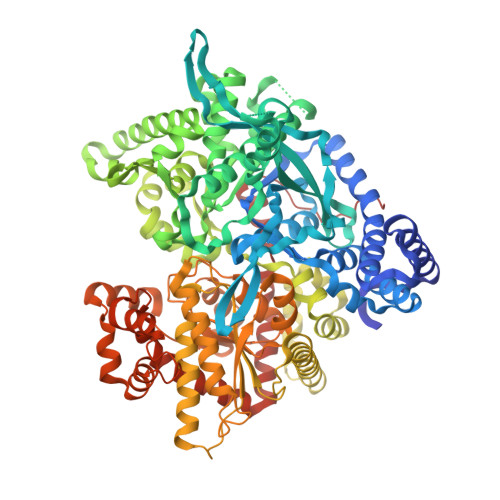
The 1.76 A Resolution Crystal Structure of Glycogen Phosphorylase B Complexed with Glucose, and Cp320626, a Potential Antidiabetic Drug
Oikonomakos, N.G., Zographos, S.E., Skamnaki, V.T., Archontis, G.(2002) Bioorg Med Chem 10: 1313
- PubMed: 11886794
- DOI: https://doi.org/10.1016/s0968-0896(01)00394-7
- Primary Citation of Related Structures:
1H5U - PubMed Abstract:
CP320626, a potential antidiabetic drug, inhibits glycogen phosphorylase in synergism with glucose. To elucidate the structural basis of synergistic inhibition, we determined the structure of muscle glycogen phosphorylase b (MGPb) complexed with both glucose and CP320626 at 1.76 A resolution, and refined to a crystallographic R value of 0.211 (R(free)=0.235). CP320626 binds at a novel allosteric site, which is some 33 A from the catalytic site, where glucose binds. The high resolution structure allows unambiguous definition of the conformation of the 1-acetyl-4-hydroxy-piperidine ring supported by theoretical energy calculations. Both CP320626 and glucose promote the less active T-state, thereby explaining their synergistic inhibition. Structural comparison of MGPb--glucose--CP320626 complex with liver glycogen phosphorylase a (LGPa) complexed with a related compound (CP403700) show that the ligand binding site is conserved in LGPa.
Organizational Affiliation:
Institute of Biological Research and Biotechnology, The National Hellenic Research Foundation, 48 Vas. Constantinou Avenue, Athens 11635, Greece. ngo@eie.gr