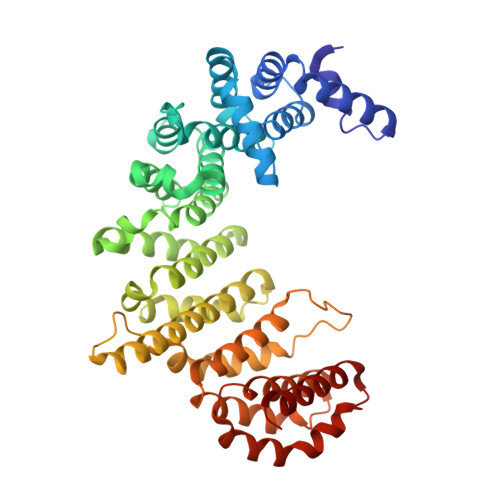
Structural basis for the interaction between FxFG nucleoporin repeats and importin-beta in nuclear trafficking.
Bayliss, R., Littlewood, T., Stewart, M.(2000) Cell 102: 99-108
- PubMed: 10929717
- DOI: https://doi.org/10.1016/s0092-8674(00)00014-3
- Primary Citation of Related Structures:
1F59 - PubMed Abstract:
We describe the crystal structure of a complex between importin-beta residues 1-442 (Ib442) and five FxFG nucleoporin repeats from Nsp1p. Nucleoporin FxFG cores bind on the convex face of Ib442 to a primary site between the A helices of HEAT repeats 5 and 6, and to a secondary site between HEAT repeats 6 and 7. Mutations at importin-beta Ile178 in the primary FxFG binding site reduce both binding and nuclear protein import, providing direct evidence for the functional significance of the importin-beta-FxFG interaction. The FxFG binding sites on importin-beta do not overlap with the RanGTP binding site. Instead, RanGTP may release importin-beta from FxFG nucleoporins by generating a conformational change that alters the structure of the FxFG binding site.
Organizational Affiliation:
MRC Laboratory of Molecular Biology, Cambridge, United Kingdom.