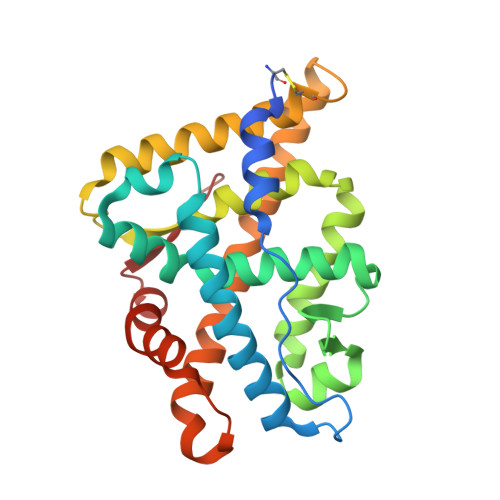
Structural evidence for ligand specificity in the binding domain of the human androgen receptor. Implications for pathogenic gene mutations.
Matias, P.M., Donner, P., Coelho, R., Thomaz, M., Peixoto, C., Macedo, S., Otto, N., Joschko, S., Scholz, P., Wegg, A., Basler, S., Schafer, M., Egner, U., Carrondo, M.A.(2000) J Biol Chem 275: 26164-26171
- PubMed: 10840043
- DOI: https://doi.org/10.1074/jbc.M004571200
- Primary Citation of Related Structures:
1E3G, 1E3K - PubMed Abstract:
The crystal structures of the human androgen receptor (hAR) and human progesterone receptor ligand-binding domains in complex with the same ligand metribolone (R1881) have been determined. Both three-dimensional structures show the typical nuclear receptor fold. The change of two residues in the ligand-binding pocket between the human progesterone receptor and hAR is most likely the source for the specificity of R1881 to the hAR. The structural implications of the 14 known mutations in the ligand-binding pocket of the hAR ligand-binding domains associated with either prostate cancer or the partial or complete androgen receptor insensitivity syndrome were analyzed. The effects of most of these mutants could be explained on the basis of the crystal structure.
Organizational Affiliation:
Instituto de Tecnologia Quimica e Biológica, Universidade Nova de Lisboa, Apartado 127, 2780 Oeiras, Portugal.