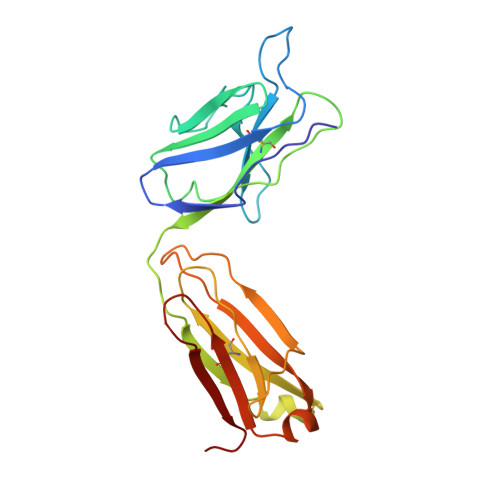
Primary and tertiary structures of the Fab fragment of a monoclonal anti-E-selectin 7A9 antibody that inhibits neutrophil attachment to endothelial cells.
Rodriguez-Romero, A., Almog, O., Tordova, M., Randhawa, Z., Gilliland, G.L.(1998) J Biol Chem 273: 11770-11775
- PubMed: 9565600
- DOI: https://doi.org/10.1074/jbc.273.19.11770
- Primary Citation of Related Structures:
1A5F - PubMed Abstract:
The murine monoclonal IgG1 antibody 7A9 binds specifically to the endothelial leukocyte adhesion molecule-1 (E-selectin), inhibiting the attachment of neutrophils to endothelial cells. The primary and three-dimensional structures of the Fab fragment of 7A9 are reported. The amino acid sequence was determined by automated Edman degradation analysis of proteolytic fragments of both the heavy and light chains of the Fab. The sequences of the two chains are consistent with that of the IgG1 class with an associated kappa light chain with two intrachain disulfide bridges in each of the heavy and light chains. The tertiary structure of the antibody fragment was determined by x-ray crystallographic methods at 2.8 A resolution. The F(ab')2 molecule, treated with dithiothreitol, crystallizes in the space group P2(1) 2(1) 2(1) with unit cell parameters a = 44.5 A, b = 83.8 A, and c = 132.5 A with one Fab molecule in the asymmetric unit. The structure was solved by the molecular replacement method and subsequently refined using simulated annealing followed by conventional least squares optimization of the coordinates. The resulting model has reasonable stereochemistry with an R factor of 0.195. The 7A9 Fab structure has an elbow bend of 162 degrees and is remarkably similar to that of the monoclonal anti-intercellular adhesion molecule-1 (ICAM-1) antibody Fab fragment. The 7A9 antigen combining site presents a groove resembling the structure of the anti-ICAM-1 antibody, and other antibodies raised against surface receptors and peptides. Residues from the six complementary determining regions (CDRs) and framework residues form the floor and walls of the groove that is approximately 22 A wide and 8 A deep and that is lined with many aromatic residues. The groove is large enough to accommodate the loop between beta-strands beta4 and beta5 of the lectin domain of E-selectin that has been implicated in neutrophil adhesion (1).
Organizational Affiliation:
Center for Advanced Research in Biotechnology of the University of Maryland Biotechnology Institute and the National Institute of Standards and Technology, Rockville, Maryland 20850, USA.