Rapid and efficient room-temperature serial synchrotron crystallography using the CFEL TapeDrive.
Zielinski, K.A., Prester, A., Andaleeb, H., Bui, S., Yefanov, O., Catapano, L., Henkel, A., Wiedorn, M.O., Lorbeer, O., Crosas, E., Meyer, J., Mariani, V., Domaracky, M., White, T.A., Fleckenstein, H., Sarrou, I., Werner, N., Betzel, C., Rohde, H., Aepfelbacher, M., Chapman, H.N., Perbandt, M., Steiner, R.A., Oberthuer, D.(2022) IUCrJ 9: 778-791
- PubMed: 36381150
- DOI: https://doi.org/10.1107/S2052252522010193
- Primary Citation of Related Structures:
7QAR, 7ZPV, 7ZQ0, 8AF4, 8AF5, 8AF6, 8AF7, 8AF8 - PubMed Abstract:
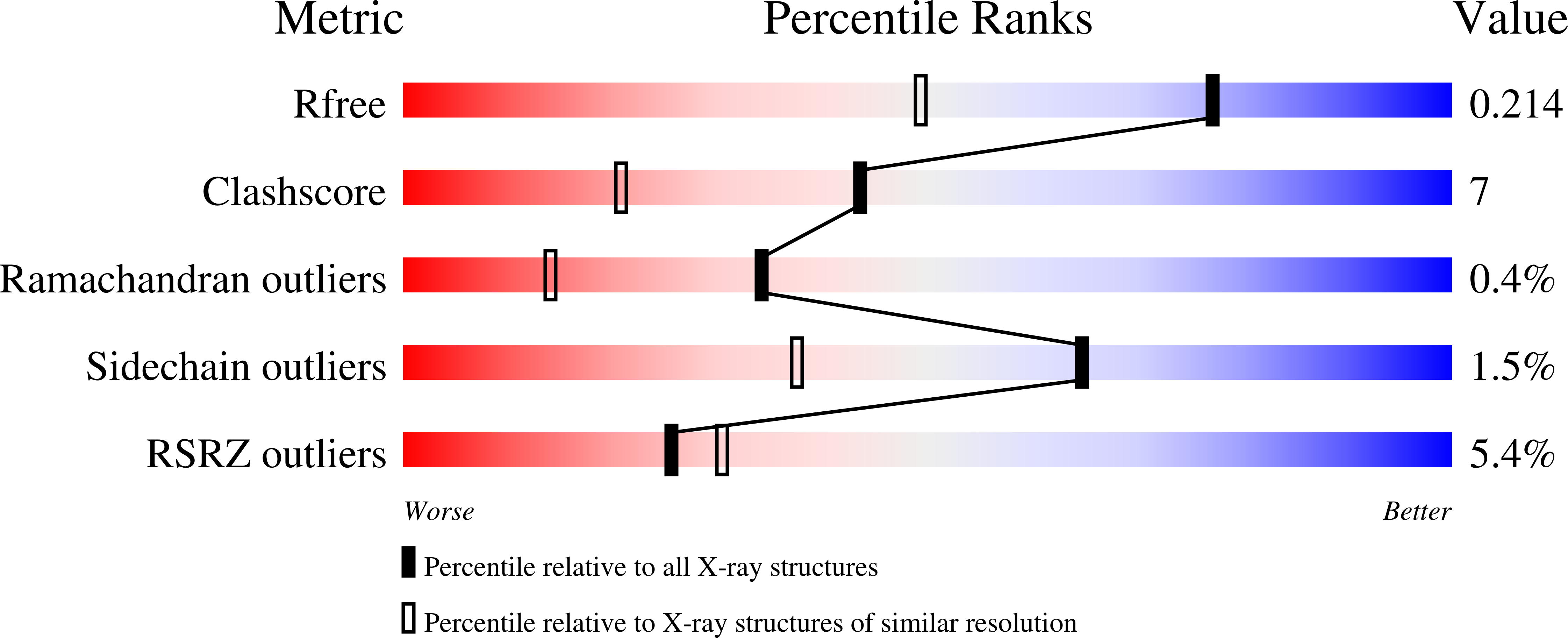
Serial crystallography at conventional synchrotron light sources (SSX) offers the possibility to routinely collect data at room temperature using micrometre-sized crystals of biological macromolecules. However, SSX data collection is not yet as routine and currently takes significantly longer than the standard rotation series cryo-crystallography. Thus, its use for high-throughput approaches, such as fragment-based drug screening, where the possibility to measure at physio-logical temperatures would be a great benefit, is impaired. On the way to high-throughput SSX using a conveyor belt based sample delivery system - the CFEL TapeDrive - with three different proteins of biological relevance ( Klebsiella pneumoniae CTX-M-14 β-lactamase, Nectria haematococca xylanase GH11 and Aspergillus flavus urate oxidase), it is shown here that complete datasets can be collected in less than a minute and only minimal amounts of sample are required.
Organizational Affiliation:
Center for Free-Electron Laser Science CFEL, Deutsches Elektronen-Synchrotron DESY, Notkestr. 85, 22607 Hamburg, Germany.