Virological characteristics of the SARS-CoV-2 Omicron BA.2 subvariants, including BA.4 and BA.5.
Kimura, I., Yamasoba, D., Tamura, T., Nao, N., Suzuki, T., Oda, Y., Mitoma, S., Ito, J., Nasser, H., Zahradnik, J., Uriu, K., Fujita, S., Kosugi, Y., Wang, L., Tsuda, M., Kishimoto, M., Ito, H., Suzuki, R., Shimizu, R., Begum, M.M., Yoshimatsu, K., Kimura, K.T., Sasaki, J., Sasaki-Tabata, K., Yamamoto, Y., Nagamoto, T., Kanamune, J., Kobiyama, K., Asakura, H., Nagashima, M., Sadamasu, K., Yoshimura, K., Shirakawa, K., Takaori-Kondo, A., Kuramochi, J., Schreiber, G., Ishii, K.J., Hashiguchi, T., Ikeda, T., Saito, A., Fukuhara, T., Tanaka, S., Matsuno, K., Sato, K.(2022) Cell 185: 3992
- PubMed: 36198317
- DOI: https://doi.org/10.1016/j.cell.2022.09.018
- Primary Citation of Related Structures:
7XWA - PubMed Abstract:
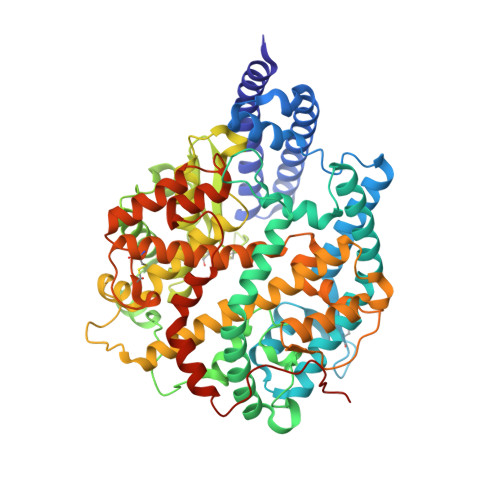
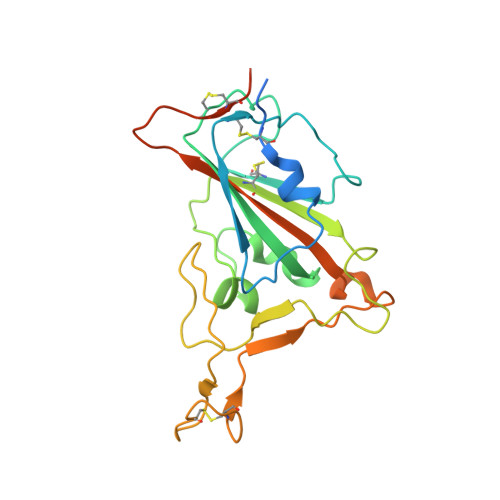
After the global spread of the SARS-CoV-2 Omicron BA.2, some BA.2 subvariants, including BA.2.9.1, BA.2.11, BA.2.12.1, BA.4, and BA.5, emerged in multiple countries. Our statistical analysis showed that the effective reproduction numbers of these BA.2 subvariants are greater than that of the original BA.2. Neutralization experiments revealed that the immunity induced by BA.1/2 infections is less effective against BA.4/5. Cell culture experiments showed that BA.2.12.1 and BA.4/5 replicate more efficiently in human alveolar epithelial cells than BA.2, and particularly, BA.4/5 is more fusogenic than BA.2. We further provided the structure of the BA.4/5 spike receptor-binding domain that binds to human ACE2 and considered how the substitutions in the BA.4/5 spike play roles in ACE2 binding and immune evasion. Moreover, experiments using hamsters suggested that BA.4/5 is more pathogenic than BA.2. Our multiscale investigations suggest that the risk of BA.2 subvariants, particularly BA.4/5, to global health is greater than that of original BA.2.
Organizational Affiliation:
Division of Systems Virology, Department of Microbiology and Immunology, The Institute of Medical Science, The University of Tokyo, Tokyo, Japan.