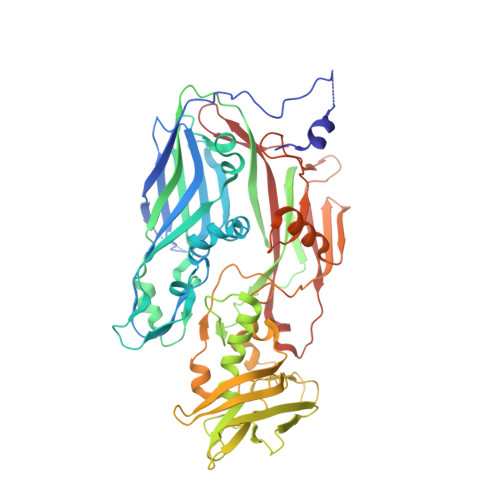
Assembly mechanism of the pleomorphic immature poxvirus scaffold.
Hyun, J., Matsunami, H., Kim, T.G., Wolf, M.(2022) Nat Commun 13: 1704-1704
- PubMed: 35361762
- DOI: https://doi.org/10.1038/s41467-022-29305-5
- Primary Citation of Related Structures:
7VFD, 7VFE, 7VFF, 7VFG, 7VFH - PubMed Abstract:
In Vaccinia virus (VACV), the prototype poxvirus, scaffold protein D13 forms a honeycomb-like lattice on the viral membrane that results in formation of the pleomorphic immature virion (IV). The structure of D13 is similar to those of major capsid proteins that readily form icosahedral capsids in nucleocytoplasmic large DNA viruses (NCLDVs). However, the detailed assembly mechanism of the nonicosahedral poxvirus scaffold has never been understood. Here we show the cryo-EM structures of the D13 trimer and scaffold intermediates produced in vitro. The structures reveal that the displacement of the short N-terminal α-helix is critical for initiation of D13 self-assembly. The continuous curvature of the IV is mediated by electrostatic interactions that induce torsion between trimers. The assembly mechanism explains the semiordered capsid-like arrangement of D13 that is distinct from icosahedral NCLDVs. Our structures explain how a single protein can self-assemble into different capsid morphologies and represent a local exception to the universal Caspar-Klug theory of quasi-equivalence.
Organizational Affiliation:
Molecular Cryo-Electron Microscopy Unit, Okinawa Institute of Science and Technology Graduate University, 1919-1 Tancha, 904-0495, Onna-son, Okinawa, Japan. jhyu002@pusan.ac.kr.