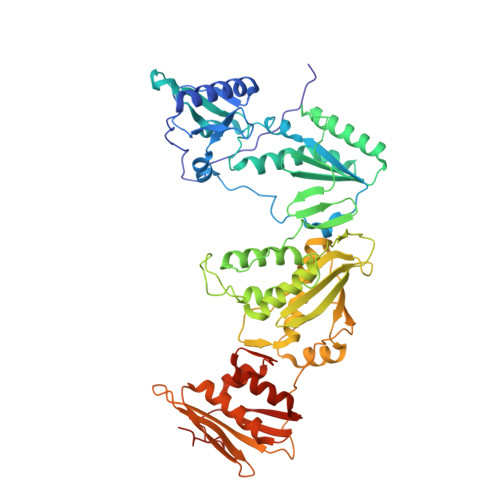
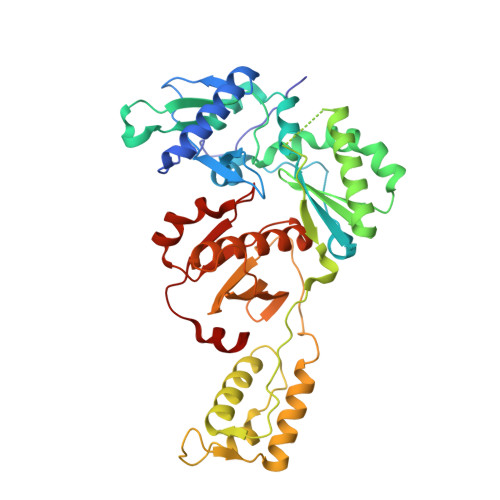
Structural Studies and Structure Activity Relationships for Novel Computationally Designed Non-nucleoside Inhibitors and Their Interactions With HIV-1 Reverse Transcriptase.
Frey, K.M., Bertoletti, N., Chan, A.H., Ippolito, J.A., Bollini, M., Spasov, K.A., Jorgensen, W.L., Anderson, K.S.(2022) Front Mol Biosci 9: 805187-805187
- PubMed: 35237658
- DOI: https://doi.org/10.3389/fmolb.2022.805187
- Primary Citation of Related Structures:
7SNP, 7SNZ, 7SO1, 7SO2, 7SO3, 7SO4, 7SO6 - PubMed Abstract:
Reverse transcriptase (RT) from the human immunodeficiency virus continues to be an attractive drug target for antiretroviral therapy. June 2022 will commemorate the 30th anniversary of the first Human Immunodeficiency Virus (HIV) RT crystal structure complex that was solved with non-nucleoside reverse transcriptase inhibitor nevirapine. The release of this structure opened opportunities for designing many families of non-nucleoside reverse transcriptase inhibitors (NNRTIs). In paying tribute to the first RT-nevirapine structure, we have developed several compound classes targeting the non-nucleoside inhibitor binding pocket of HIV RT. Extensive analysis of crystal structures of RT in complex with the compounds informed iterations of structure-based drug design. Structures of seven additional complexes were determined and analyzed to summarize key interactions with residues in the non-nucleoside inhibitor binding pocket (NNIBP) of RT. Additional insights comparing structures with antiviral data and results from molecular dynamics simulations elucidate key interactions and dynamics between the nucleotide and non-nucleoside binding sites.
Organizational Affiliation:
Department of Pharmacology, Yale University School of Medicine, New Haven, CT, United States.