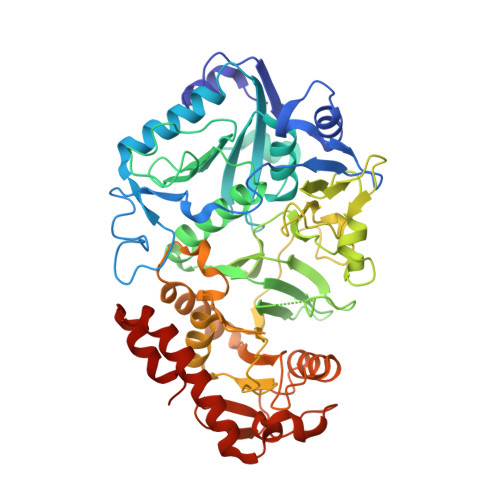
Kinetic and structural analysis of Escherichia coli phosphoenolpyruvate carboxykinase mutants.
Sokaribo, A., Novakovski, B.A.A., Cotelesage, J., White, A.P., Sanders, D., Goldie, H.(2020) Biochim Biophys Acta Gen Subj 1864: 129517-129517
- PubMed: 31911238
- DOI: https://doi.org/10.1016/j.bbagen.2020.129517
- Primary Citation of Related Structures:
6COM, 6V2L, 6V2M, 6V2N - PubMed Abstract:
Phosphoenolpyruvate carboxykinase (PEPCK) is a metabolic enzyme in the gluconeogenesis pathway, where it catalyzes the reversible conversion of oxaloacetate (OAA) to phosphoenolpyruvate (PEP) and CO 2 . The substrates for Escherichia coli PEPCK are OAA and MgATP, with Mn 2+ acting as a cofactor. Analysis of PEPCK structures have revealed amino acid residues involved in substrate/cofactor coordination during catalysis.
Organizational Affiliation:
Department of Biochemistry, Microbiology and Immunology, University of Saskatchewan, Saskatoon, SK, Canada; Vaccine and Infectious Disease Organization, International Vaccine Center, Saskatoon, SK, Canada. Electronic address: ats172@mail.usask.ca.