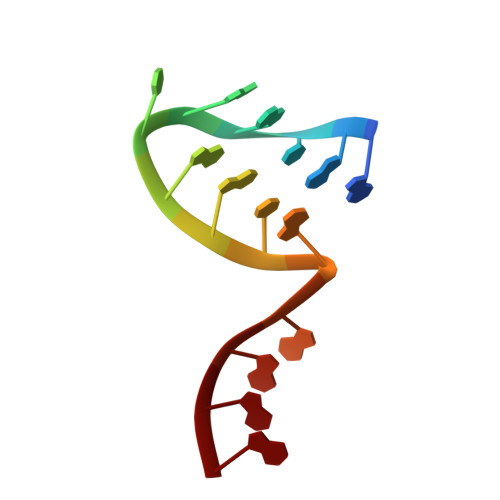
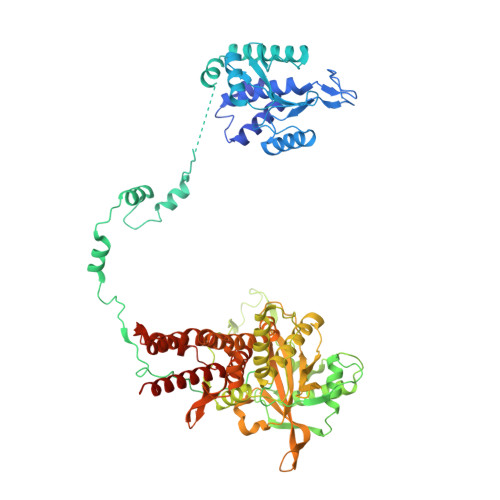
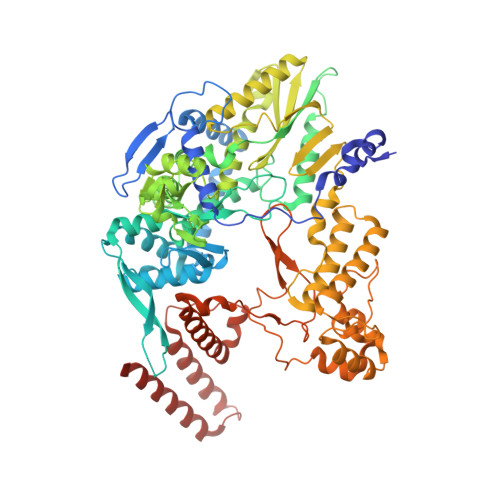
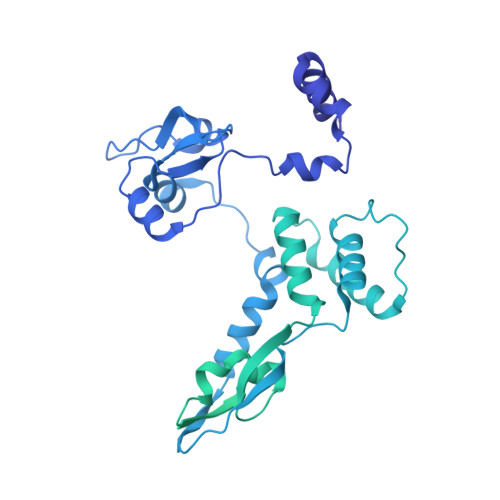
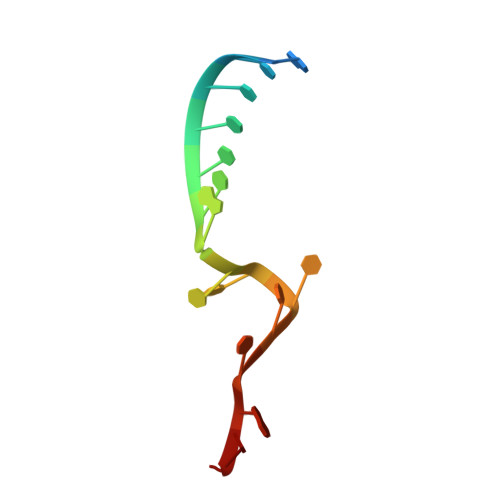
A Structure-Based Model for the Complete Transcription Cycle of Influenza Polymerase.
Wandzik, J.M., Kouba, T., Karuppasamy, M., Pflug, A., Drncova, P., Provaznik, J., Azevedo, N., Cusack, S.(2020) Cell 181: 877
- PubMed: 32304664
- DOI: https://doi.org/10.1016/j.cell.2020.03.061
- Primary Citation of Related Structures:
6SZU, 6SZV, 6T0N, 6T0R, 6T0S, 6T0U, 6T0V, 6T0W, 6T2C, 6TU5, 6TW1 - PubMed Abstract:
Influenza polymerase uses unique mechanisms to synthesize capped and polyadenylated mRNAs from the genomic viral RNA (vRNA) template, which is packaged inside ribonucleoprotein particles (vRNPs). Here, we visualize by cryoelectron microscopy the conformational dynamics of the polymerase during the complete transcription cycle from pre-initiation to termination, focusing on the template trajectory. After exiting the active site cavity, the template 3' extremity rebinds into a specific site on the polymerase surface. Here, it remains sequestered during all subsequent transcription steps, forcing the template to loop out as it further translocates. At termination, the strained connection between the bound template 5' end and the active site results in polyadenylation by stuttering at uridine 17. Upon product dissociation, further conformational changes release the trapped template, allowing recycling back into the pre-initiation state. Influenza polymerase thus performs transcription while tightly binding to and protecting both template ends, allowing efficient production of multiple mRNAs from a single vRNP.
Organizational Affiliation:
European Molecular Biology Laboratory, 71 Avenue des Martyrs, CS 90181, 38042 Grenoble Cedex 9, France.