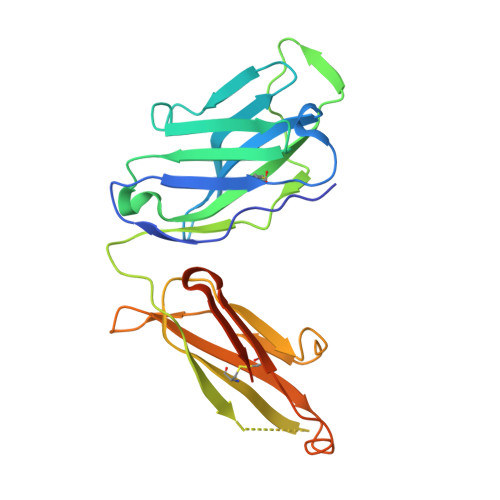
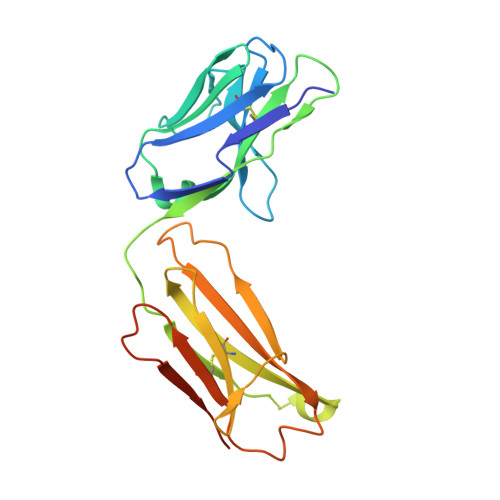
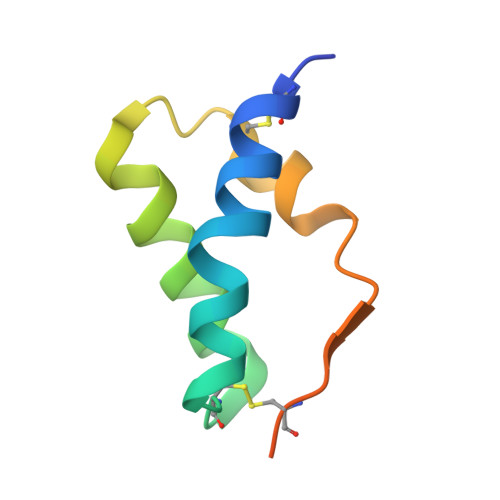
De novo protein design enables the precise induction of RSV-neutralizing antibodies.
Sesterhenn, F., Yang, C., Bonet, J., Cramer, J.T., Wen, X., Wang, Y., Chiang, C.I., Abriata, L.A., Kucharska, I., Castoro, G., Vollers, S.S., Galloux, M., Dheilly, E., Rosset, S., Corthesy, P., Georgeon, S., Villard, M., Richard, C.A., Descamps, D., Delgado, T., Oricchio, E., Rameix-Welti, M.A., Mas, V., Ervin, S., Eleouet, J.F., Riffault, S., Bates, J.T., Julien, J.P., Li, Y., Jardetzky, T., Krey, T., Correia, B.E.(2020) Science 368
- PubMed: 32409444
- DOI: https://doi.org/10.1126/science.aay5051
- Primary Citation of Related Structures:
6S3D, 6VTW, 6XWI, 6XXV - PubMed Abstract:
De novo protein design has been successful in expanding the natural protein repertoire. However, most de novo proteins lack biological function, presenting a major methodological challenge. In vaccinology, the induction of precise antibody responses remains a cornerstone for next-generation vaccines. Here, we present a protein design algorithm called TopoBuilder, with which we engineered epitope-focused immunogens displaying complex structural motifs. In both mice and nonhuman primates, cocktails of three de novo-designed immunogens induced robust neutralizing responses against the respiratory syncytial virus. Furthermore, the immunogens refocused preexisting antibody responses toward defined neutralization epitopes. Overall, our design approach opens the possibility of targeting specific epitopes for the development of vaccines and therapeutic antibodies and, more generally, will be applicable to the design of de novo proteins displaying complex functional motifs.
Organizational Affiliation:
Institute of Bioengineering, École Polytechnique Fédérale de Lausanne, Lausanne CH-1015, Switzerland.