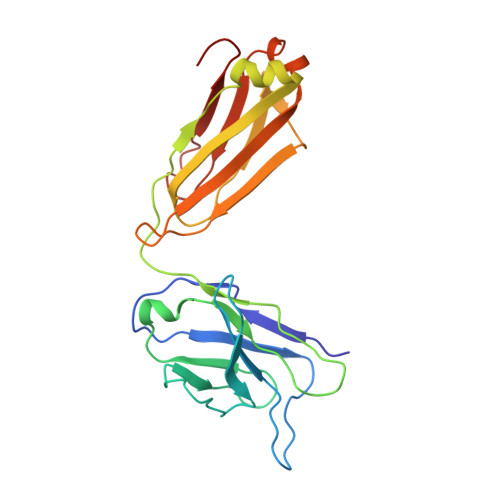
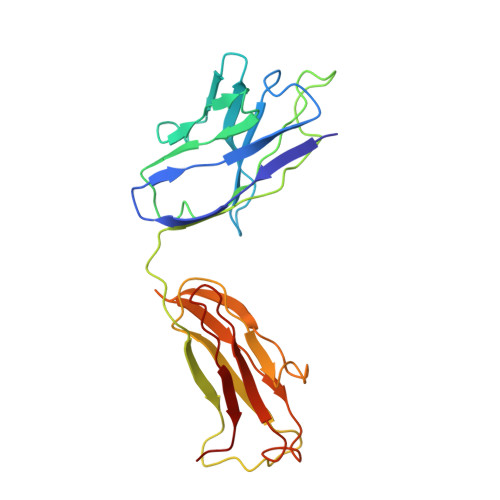
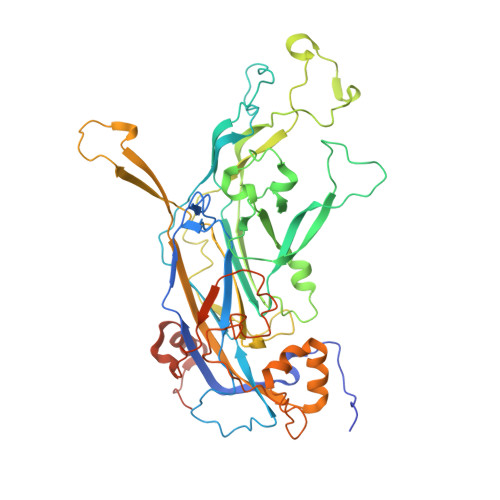
High-Resolution Structure Analysis of Antibody V5 and U4 Conformational Epitopes on Human Papillomavirus 16.
Guan, J., Bywaters, S.M., Brendle, S.A., Ashley, R.E., Makhov, A.M., Conway, J.F., Christensen, N.D., Hafenstein, S.(2017) Viruses 9
- PubMed: 29211035
- DOI: https://doi.org/10.3390/v9120374
- Primary Citation of Related Structures:
6BSP, 6BT3 - PubMed Abstract:
Cancers attributable to human papillomavirus (HPV) place a huge burden on the health of both men and women. The current commercial vaccines are genotype specific and provide little therapeutic benefit to patients with existing HPV infections. Identifying the conformational epitopes on the virus capsid supports the development of improved recombinant vaccines to maximize long-term protection against multiple types of HPV. Fragments of antibody (Fab) digested from the neutralizing monoclonal antibodies H16.V5 (V5) and H16.U4 (U4) were bound to HPV16 capsids and the structures of the two virus-Fab complexes were solved to near atomic resolution using cryo-electron microscopy. The structures reveal virus conformational changes, the Fab-binding mode to the capsid, the residues comprising the epitope and indicate a potential interaction of U4 with the minor structural protein, L2. Competition enzyme-linked immunosorbent assay (ELISA) showed V5 outcompetes U4 when added sequentially, demonstrating a steric interference even though the footprints do not overlap. Combined with our previously reported immunological and structural results, we propose that the virus may initiate host entry through an interaction between the icosahedral five-fold vertex of the capsid and receptors on the host cell. The highly detailed epitopes identified for the two antibodies provide a framework for continuing biochemical, genetic and biophysical studies.
Organizational Affiliation:
Department of Medicine, The Pennsylvania State University College of Medicine, 500 University Drive, Hershey, PA 17033, USA. fuyougj@gmail.com.