Crystal structure of tryptophan synthase from M. tuberculosis - open form with BRD6309 bound
Chang, C., Michalska, K., Maltseva, N.I., Jedrzejczak, R., McCarren, P., Nag, P.P., Joachimiak, A., Satchell, K.To be published.
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Tryptophan synthase alpha chain | 276 | Mycobacterium tuberculosis H37Rv | Mutation(s): 0 Gene Names: trpA, Rv1613, MTCY01B2.05 EC: 4.2.1.20 | 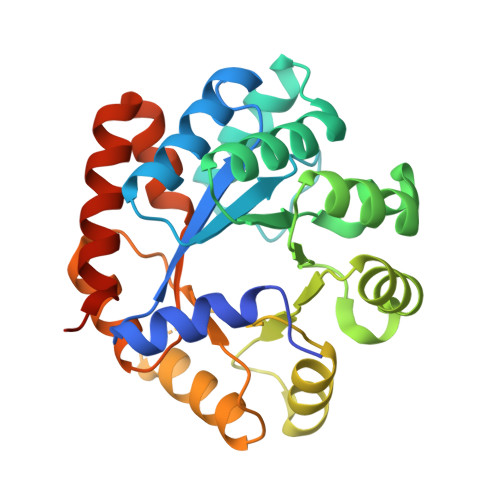 | |
UniProt | |||||
Find proteins for P9WFY1 (Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv)) Explore P9WFY1 Go to UniProtKB: P9WFY1 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P9WFY1 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Tryptophan synthase beta chain | 410 | Mycobacterium tuberculosis H37Rv | Mutation(s): 0 Gene Names: trpB, Rv1612, MTCY01B2.04 EC: 4.2.1.20 | 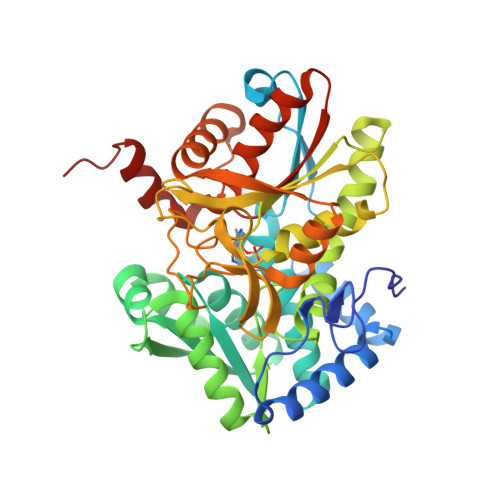 | |
UniProt | |||||
Find proteins for P9WFX9 (Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv)) Explore P9WFX9 Go to UniProtKB: P9WFX9 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P9WFX9 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 6 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| H9V (Subject of Investigation/LOI) Query on H9V | KA [auth F], L [auth B], UA [auth H], W [auth D] | (2R,3S,4R)-3-(4'-chloro-2',6'-difluoro[1,1'-biphenyl]-4-yl)-4-(fluoromethyl)azetidine-2-carbonitrile C17 H12 Cl F3 N2 XHUZEPHBFHVYMY-YQQAZPJKSA-N |  | ||
| MLT Query on MLT | FA [auth D], GA [auth D], P [auth B], QA [auth F], WA [auth H] | D-MALATE C4 H6 O5 BJEPYKJPYRNKOW-UWTATZPHSA-N |  | ||
| MLA Query on MLA | O [auth B], PA [auth F], U [auth C] | MALONIC ACID C3 H4 O4 OFOBLEOULBTSOW-UHFFFAOYSA-N |  | ||
| EDO Query on EDO | HA [auth D], K [auth A] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| ACT Query on ACT | IA [auth D] JA [auth D] TA [auth G] X [auth D] XA [auth H] | ACETATE ION C2 H3 O2 QTBSBXVTEAMEQO-UHFFFAOYSA-M |  | ||
| FMT Query on FMT | AA [auth D] BA [auth D] CA [auth D] DA [auth D] EA [auth D] | FORMIC ACID C H2 O2 BDAGIHXWWSANSR-UHFFFAOYSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| LLP Query on LLP | B, D, F, H | L-PEPTIDE LINKING | C14 H22 N3 O7 P |  | LYS |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 134.777 | α = 90 |
| b = 157.877 | β = 90 |
| c = 166.69 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| SCALEPACK | data scaling |
| PDB_EXTRACT | data extraction |
| HKL-3000 | data reduction |
| HKL-3000 | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Structure-guided Drug Discovery Coalition | Canada | -- |