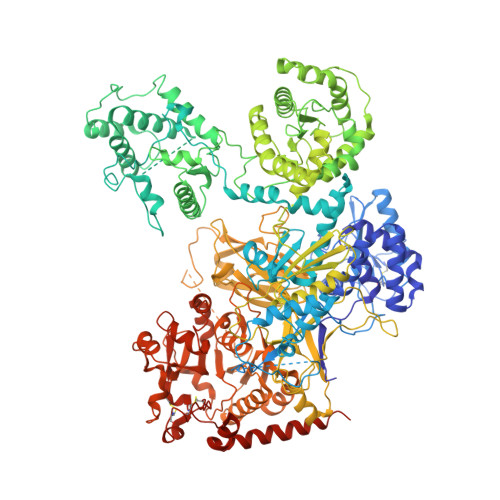
Clamping, bending, and twisting inter-domain motions in the misfold-recognizing portion of UDP-glucose: Glycoprotein glucosyltransferase.
Modenutti, C.P., Blanco Capurro, J.I., Ibba, R., Alonzi, D.S., Song, M.N., Vasiljevic, S., Kumar, A., Chandran, A.V., Tax, G., Marti, L., Hill, J.C., Lia, A., Hensen, M., Waksman, T., Rushton, J., Rubichi, S., Santino, A., Marti, M.A., Zitzmann, N., Roversi, P.(2021) Structure 29: 357
- PubMed: 33352114
- DOI: https://doi.org/10.1016/j.str.2020.11.017
- Primary Citation of Related Structures:
6TRF, 6TRT, 6TS2, 6TS8 - PubMed Abstract:
UDP-glucose:glycoprotein glucosyltransferase (UGGT) flags misfolded glycoproteins for ER retention. We report crystal structures of full-length Chaetomium thermophilum UGGT (CtUGGT), two CtUGGT double-cysteine mutants, and its TRXL2 domain truncation (CtUGGT-ΔTRXL2). CtUGGT molecular dynamics (MD) simulations capture extended conformations and reveal clamping, bending, and twisting inter-domain movements. We name "Parodi limit" the maximum distance on the same glycoprotein between a site of misfolding and an N-linked glycan that can be reglucosylated by monomeric UGGT in vitro, in response to recognition of misfold at that site. Based on the MD simulations, we estimate the Parodi limit as around 70-80 Å. Frequency distributions of distances between glycoprotein residues and their closest N-linked glycosylation sites in glycoprotein crystal structures suggests relevance of the Parodi limit to UGGT activity in vivo. Our data support a "one-size-fits-all adjustable spanner" UGGT substrate recognition model, with an essential role for the UGGT TRXL2 domain.
Organizational Affiliation:
Departamento de Química Biológica, Facultad de Ciencias Exactas y Naturales, Universidad de Buenos Aires, Ciudad Universitaria, Pab. II (CE1428EHA), Buenos Aires, Argentina; Instituto de Química Biológica de la Facultad de Ciencias Exactas y Naturales (IQUIBICEN) CONICET. Ciudad Universitaria, Pab. II (CE1428EHA), Buenos Aires, Argentina.