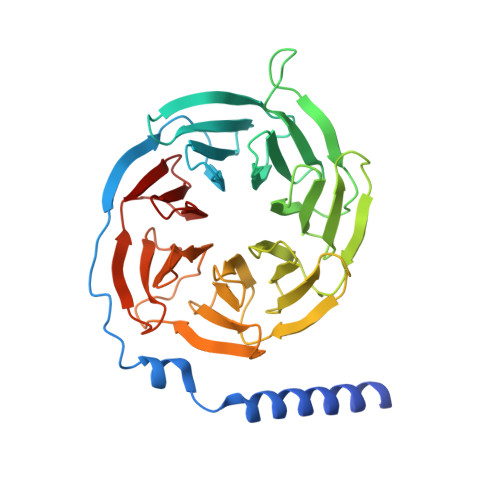
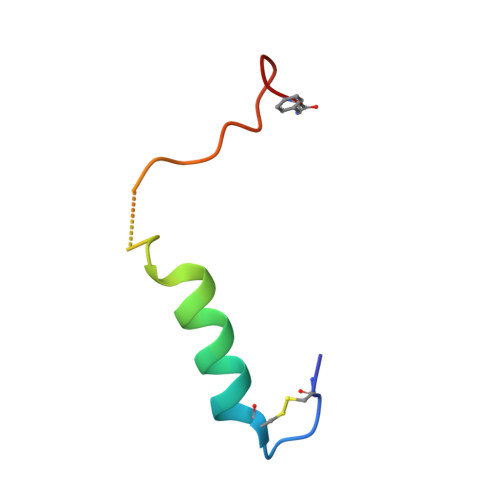
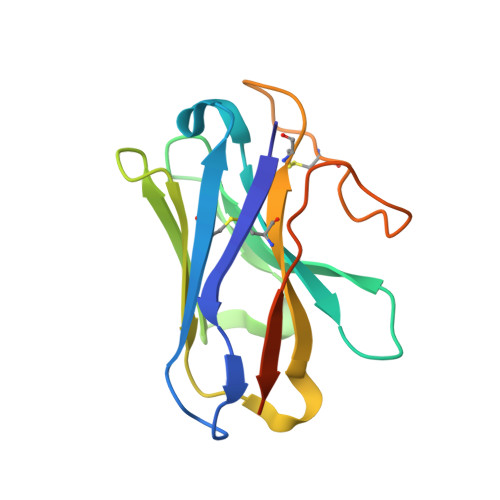
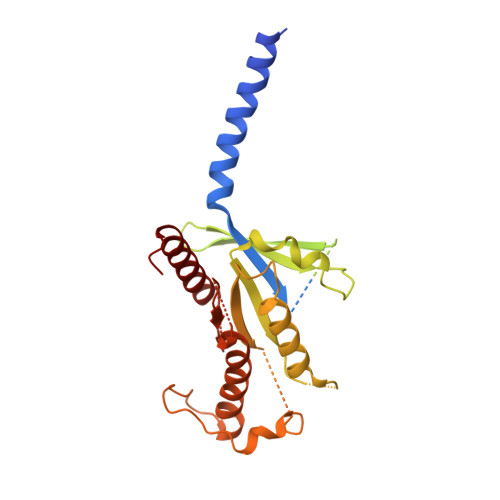
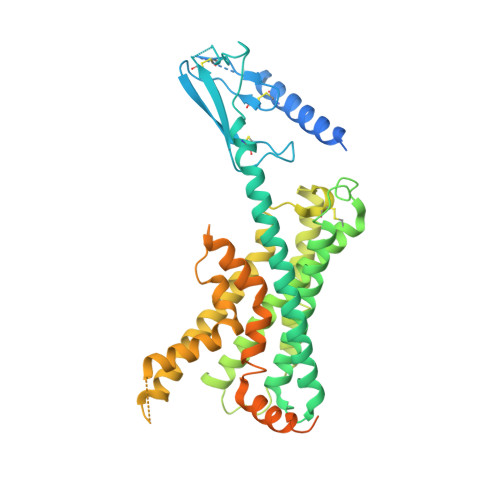
Cryo-EM structure of the active, Gs-protein complexed, human CGRP receptor.
Liang, Y.L., Khoshouei, M., Deganutti, G., Glukhova, A., Koole, C., Peat, T.S., Radjainia, M., Plitzko, J.M., Baumeister, W., Miller, L.J., Hay, D.L., Christopoulos, A., Reynolds, C.A., Wootten, D., Sexton, P.M.(2018) Nature 561: 492-497
- PubMed: 30209400
- DOI: https://doi.org/10.1038/s41586-018-0535-y
- Primary Citation of Related Structures:
6E3Y - PubMed Abstract:
Calcitonin gene-related peptide (CGRP) is a widely expressed neuropeptide that has a major role in sensory neurotransmission. The CGRP receptor is a heterodimer of the calcitonin receptor-like receptor (CLR) class B G-protein-coupled receptor and a type 1 transmembrane domain protein, receptor activity-modifying protein 1 (RAMP1). Here we report the structure of the human CGRP receptor in complex with CGRP and the G s -protein heterotrimer at 3.3 Å global resolution, determined by Volta phase-plate cryo-electron microscopy. The receptor activity-modifying protein transmembrane domain sits at the interface between transmembrane domains 3, 4 and 5 of CLR, and stabilizes CLR extracellular loop 2. RAMP1 makes only limited direct contact with CGRP, consistent with its function in allosteric modulation of CLR. Molecular dynamics simulations indicate that RAMP1 provides stability to the receptor complex, particularly in the positioning of the extracellular domain of CLR. This work provides insights into the control of G-protein-coupled receptor function.
Organizational Affiliation:
Drug Discovery Biology and Department of Pharmacology, Monash Institute of Pharmaceutical Sciences, Monash University, Parkville, Victoria, Australia.