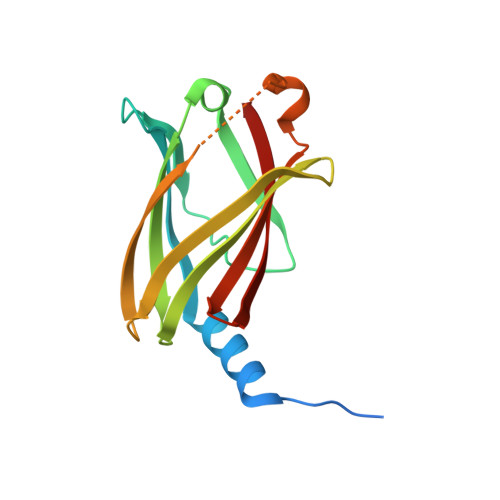
Mechanistic insights into the role of prenyl-binding protein PrBP/ delta in membrane dissociation of phosphodiesterase 6.
Qureshi, B.M., Schmidt, A., Behrmann, E., Burger, J., Mielke, T., Spahn, C.M.T., Heck, M., Scheerer, P.(2018) Nat Commun 9: 90-90
- PubMed: 29311697
- DOI: https://doi.org/10.1038/s41467-017-02569-y
- PubMed Abstract:
Isoprenylated proteins are associated with membranes and their inter-compartmental distribution is regulated by solubilization factors, which incorporate lipid moieties in hydrophobic cavities and thereby facilitate free diffusion during trafficking. Here we report the crystal structure of a solubilization factor, the prenyl-binding protein (PrBP/δ), at 1.81 Å resolution in its ligand-free apo-form. Apo-PrBP/δ harbors a preshaped, deep hydrophobic cavity, capacitating apo-PrBP/δ to readily bind its prenylated cargo. To investigate the molecular mechanism of cargo solubilization we analyzed the PrBP/δ-induced membrane dissociation of rod photoreceptor phosphodiesterase (PDE6). The results suggest that PrBP/δ exclusively interacts with the soluble fraction of PDE6. Depletion of soluble species in turn leads to dissociation of membrane-bound PDE6, as both are in equilibrium. This "solubilization by depletion" mechanism of PrBP/δ differs from the extraction of prenylated proteins by the similar folded solubilization factor RhoGDI, which interacts with membrane bound cargo via an N-terminal structural element lacking in PrBP/δ.
Organizational Affiliation:
Charité - Universitätsmedizin Berlin, corporate member of Freie Universität Berlin, Humboldt-Universität zu Berlin, and Berlin Institute of Health, Charitéplatz 1, D-10117, Berlin, Germany.