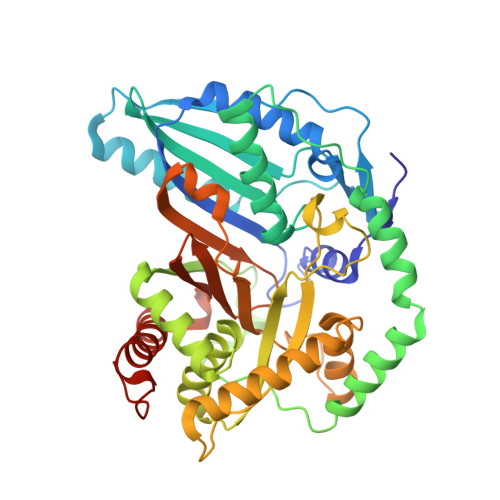
Structural and mutational analysis of the nonribosomal peptide synthetase heterocyclization domain provides insight into catalysis.
Bloudoff, K., Fage, C.D., Marahiel, M.A., Schmeing, T.M.(2017) Proc Natl Acad Sci U S A 114: 95-100
- PubMed: 27994138
- DOI: https://doi.org/10.1073/pnas.1614191114
- Primary Citation of Related Structures:
5T3E - PubMed Abstract:
Nonribosomal peptide synthetases (NRPSs) are a family of multidomain, multimodule enzymes that synthesize structurally and functionally diverse peptides, many of which are of great therapeutic or commercial value. The central chemical step of peptide synthesis is amide bond formation, which is typically catalyzed by the condensation (C) domain. In many NRPS modules, the C domain is replaced by the heterocyclization (Cy) domain, a homologous domain that performs two consecutive reactions by using hitherto unknown catalytic mechanisms. It first catalyzes amide bond formation, and then the intramolecular cyclodehydration between a Cys, Ser, or Thr side chain and the backbone carbonyl carbon to form a thiazoline, oxazoline, or methyloxazoline ring. The rings are important for the form and function of the peptide product. We present the crystal structure of an NRPS Cy domain, Cy2 of bacillamide synthetase, at a resolution of 2.3 Å. Despite sharing the same fold, the active sites of C and Cy domains have important differences. The structure allowed us to probe the roles of active-site residues by using mutational analyses in a peptide synthesis assay with intact bacillamide synthetase. The drastically different effects of these mutants, interpreted by using our structural and bioinformatic results, provide insight into the catalytic mechanisms of the Cy domain and implicate a previously unexamined Asp-Thr dyad in catalysis of the cyclodehydration reaction.
Organizational Affiliation:
Department of Biochemistry, McGill University, Montreal, QC H3G 0B1, Canada.