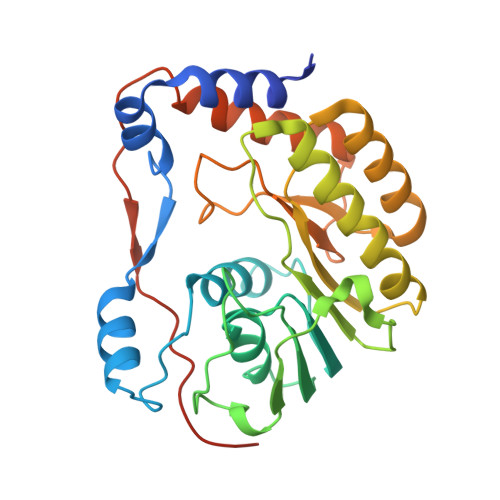
The crystal structure of Zika virus NS5 reveals conserved drug targets.
Duan, W., Song, H., Wang, H., Chai, Y., Su, C., Qi, J., Shi, Y., Gao, G.F.(2017) EMBO J 36: 919-933
- PubMed: 28254839
- DOI: https://doi.org/10.15252/embj.201696241
- Primary Citation of Related Structures:
5WZ1, 5WZ2, 5WZ3 - PubMed Abstract:
Zika virus (ZIKV) has emerged as major health concern, as ZIKV infection has been shown to be associated with microcephaly, severe neurological disease and possibly male sterility. As the largest protein component within the ZIKV replication complex, NS5 plays key roles in the life cycle and survival of the virus through its N-terminal methyltransferase (MTase) and C-terminal RNA-dependent RNA polymerase (RdRp) domains. Here, we present the crystal structures of ZIKV NS5 MTase in complex with an RNA cap analogue ( m7 GpppA) and the free NS5 RdRp. We have identified the conserved features of ZIKV NS5 MTase and RdRp structures that could lead to development of current antiviral inhibitors being used against flaviviruses, including dengue virus and West Nile virus, to treat ZIKV infection. These results should inform and accelerate the structure-based design of antiviral compounds against ZIKV.
Organizational Affiliation:
CAS Key Laboratory of Pathogenic Microbiology and Immunology, Institute of Microbiology, Chinese Academy of Sciences, Beijing, China.