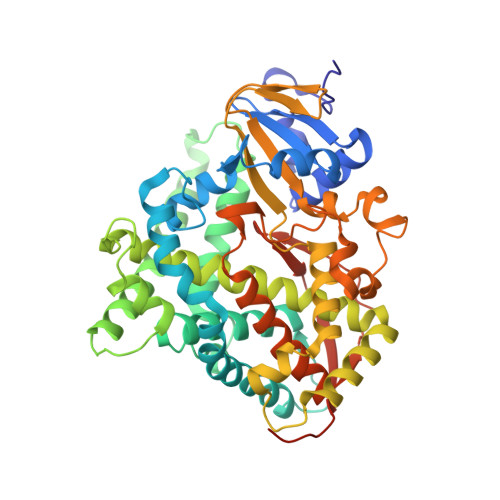
An Evolved Orthogonal Enzyme/Cofactor Pair.
Reynolds, E.W., McHenry, M.W., Cannac, F., Gober, J.G., Snow, C.D., Brustad, E.M.(2016) J Am Chem Soc 138: 12451-12458
- PubMed: 27575374
- DOI: https://doi.org/10.1021/jacs.6b05847
- Primary Citation of Related Structures:
5JQU, 5JQV - PubMed Abstract:
We introduce a strategy that expands the functionality of hemoproteins through orthogonal enzyme/heme pairs. By exploiting the ability of a natural heme transport protein, ChuA, to promiscuously import heme derivatives, we have evolved a cytochrome P450 (P450BM3) that selectively incorporates a nonproteinogenic cofactor, iron deuteroporphyrin IX (Fe-DPIX), even in the presence of endogenous heme. Crystal structures show that selectivity gains are due to mutations that introduce steric clash with the heme vinyl groups while providing a complementary binding surface for the smaller Fe-DPIX cofactor. Furthermore, the evolved orthogonal enzyme/cofactor pair is active in non-natural carbenoid-mediated olefin cyclopropanation. This methodology for the generation of orthogonal enzyme/cofactor pairs promises to expand cofactor diversity in artificial metalloenzymes.
Organizational Affiliation:
Department of Chemistry, University of North Carolina-Chapel Hill , 125 South Road, CB 3290, Chapel Hill, North Carolina 27599, United States.