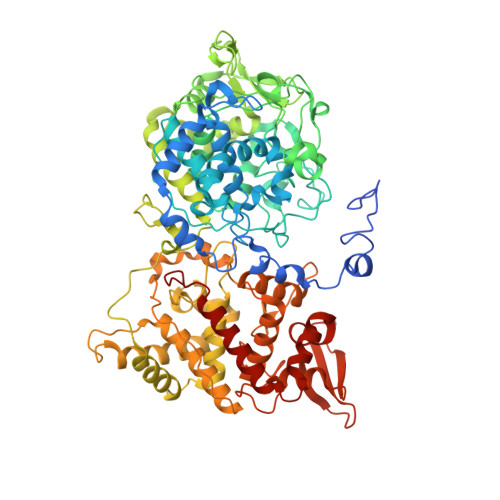
Interaction with the Redox Cofactor MYW and Functional Role of a Mobile Arginine in Eukaryotic Catalase-Peroxidase.
Gasselhuber, B., Graf, M.M., Jakopitsch, C., Zamocky, M., Nicolussi, A., Furtmuller, P.G., Oostenbrink, C., Carpena, X., Obinger, C.(2016) Biochemistry 55: 3528-3541
- PubMed: 27293030
- DOI: https://doi.org/10.1021/acs.biochem.6b00436
- Primary Citation of Related Structures:
5JHX, 5JHY, 5JHZ - PubMed Abstract:
Catalase-peroxidases (KatGs) are unique bifunctional heme peroxidases with an additional posttranslationally formed redox-active Met-Tyr-Trp cofactor that is essential for catalase activity. On the basis of studies of bacterial KatGs, controversial mechanisms of hydrogen peroxide oxidation were proposed. The recent discovery of eukaryotic KatGs with differing pH optima of catalase activity now allows us to scrutinize those postulated reaction mechanisms. In our study, secreted KatG from the fungus Magnaporthe grisea (MagKatG2) was used to analyze the role of a remote KatG-typical mobile arginine that was shown to interact with the Met-Tyr-Trp adduct in a pH-dependent manner in bacterial KatGs. Here we present crystal structures of MagKatG2 at pH 3.0, 5.5, and 7.0 and investigate the mobility of Arg461 by molecular dynamics simulation. Data suggest that at pH ≥4.5 Arg461 mostly interacts with the deprotonated adduct Tyr. Elimination of Arg461 by mutation to Ala slightly increases the thermal stability but does not alter the active site architecture or the kinetics of cyanide binding. However, the variant Arg461Ala lost the wild-type-typical optimum of catalase activity at pH 5.25 (kcat = 6450 s(-1)) but exhibits a broad plateau between pH 4.5 and 7.5 (kcat = 270 s(-1) at pH 5.5). Moreover, significant differences in the kinetics of interconversion of redox intermediates of wild-type and mutant protein mixed with either peroxyacetic acid or hydrogen peroxide are observed. These findings together with published data from bacterial KatGs allow us to propose a role of Arg461 in the H2O2 oxidation reaction of KatG.
Organizational Affiliation:
Department of Chemistry, Division of Biochemistry, BOKU-University of Natural Resources and Life Sciences , Muthgasse 18, A-1190 Vienna, Austria.