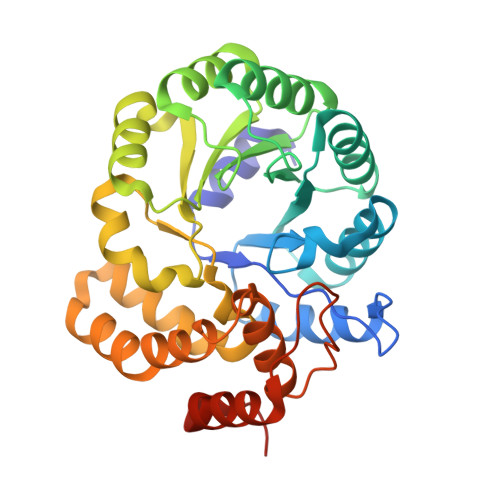
Structure and function of a decarboxylating Agrobacterium tumefaciens keto-deoxy-d-galactarate dehydratase.
Taberman, H., Andberg, M., Parkkinen, T., Janis, J., Penttila, M., Hakulinen, N., Koivula, A., Rouvinen, J.(2014) Biochemistry 53: 8052-8060
- PubMed: 25454257
- DOI: https://doi.org/10.1021/bi501290k
- Primary Citation of Related Structures:
4UR7, 4UR8, 5HWJ, 5HWM, 5HWN - PubMed Abstract:
Agrobacterium tumefaciens (At) strain C58 contains an oxidative enzyme pathway that can function on both d-glucuronic and d-galacturonic acid. The corresponding gene coding for At keto-deoxy-d-galactarate (KDG) dehydratase is located in the same gene cluster as those coding for uronate dehydrogenase (At Udh) and galactarolactone cycloisomerase (At Gci) which we have previously characterized. Here, we present the kinetic characterization and crystal structure of At KDG dehydratase, which catalyzes the next step, the decarboxylating hydrolyase reaction of KDG to produce α-ketoglutaric semialdehyde (α-KGSA) and carbon dioxide. The crystal structures of At KDG dehydratase and its complexes with pyruvate and 2-oxoadipic acid, two substrate analogues, were determined to 1.7 Å, 1.5 Å, and 2.1 Å resolution, respectively. Furthermore, mass spectrometry was used to confirm reaction end-products. The results lead us to propose a structure-based mechanism for At KDG dehydratase, suggesting that while the enzyme belongs to the Class I aldolase protein family, it does not follow a typical retro-aldol condensation mechanism.
Organizational Affiliation:
Department of Chemistry, University of Eastern Finland , FI-80101 Joensuu, Finland.