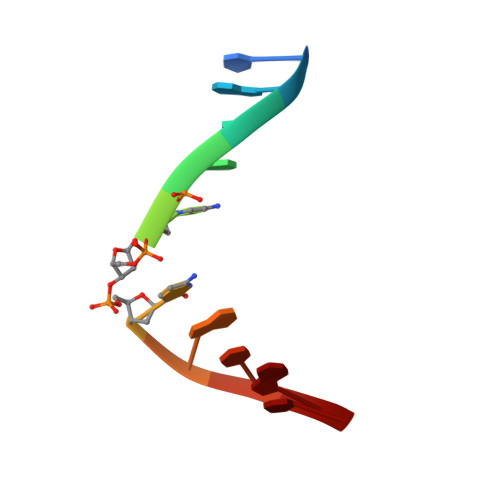
Nuclear Magnetic Resonance Solution Structure of DNA Featuring Clustered 2'-Deoxyribonolactone and 8-Oxoguanine Lesions.
Zalesak, J., Constant, J.F., Jourdan, M.(2016) Biochemistry 55: 3899-3906
- PubMed: 27322640
- DOI: https://doi.org/10.1021/acs.biochem.6b00396
- Primary Citation of Related Structures:
5HQF, 5HQQ - PubMed Abstract:
Ionizing radiation, free radicals, and reactive oxygen species produce hundreds of different DNA lesions. Clustered lesions are typical for ionizing radiation. They compromise the efficiency of the base excision repair (BER) pathway, and as a consequence, they are much more toxic and mutagenic than isolated lesions. Despite their biological relevance, e.g., in cancer radiotherapy and accidental exposure, they are not very well studied from a structural point of view, and while insights provided by structural studies contribute to the understanding of the repair process, only three nuclear magnetic resonance (NMR) studies of DNA containing clusters of lesions were reported. Herein, we report the first NMR solution structure of two DNAs containing a bistranded cluster with the 2'-deoxyribonolactone and 8-oxoguanine lesions. Both DNA duplexes feature a 2'-deoxyribonolactone site in the middle of the sequence of one strand and differ by the relative position of the 8-oxoguanine, staggered 3' or 5' side on the complementary strand at a three-nucleotide distance. Depending on its relative position, the repair of the 8-oxoguanine lesion by the base excision repair protein Fpg is either almost complete or inhibited. We found that the structures of the two DNAs containing a bistranded cluster of two lesions are similar and do not deviate very much from the standard B-form. As no obvious structural deformations were observed between the two duplexes, we concluded that the differences in Fpg activity are not due to differences in their global conformation.
Organizational Affiliation:
Universite Grenoble Alpes , DCM UMR 5250, F-38000 Grenoble, France.