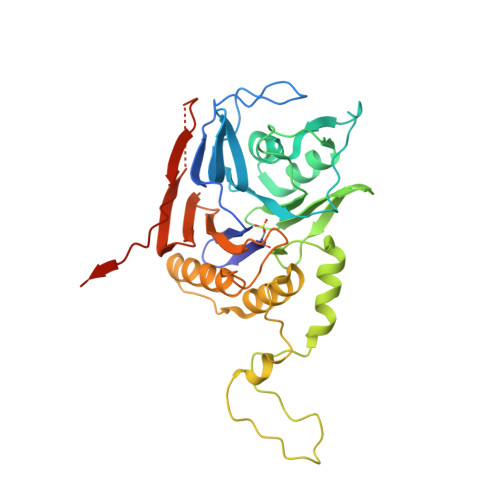
Crystal structure of bile salt hydrolase from Lactobacillus salivarius.
Xu, F., Guo, F., Hu, X.J., Lin, J.(2016) Acta Crystallogr F Struct Biol Commun 72: 376-381
- PubMed: 27139829
- DOI: https://doi.org/10.1107/S2053230X16005707
- Primary Citation of Related Structures:
5HKE - PubMed Abstract:
Bile salt hydrolase (BSH) is a gut-bacterial enzyme that negatively influences host fat digestion and energy harvesting. The BSH enzyme activity functions as a gateway reaction in the small intestine by the deconjugation of glycine-conjugated or taurine-conjugated bile acids. Extensive gut-microbiota studies have suggested that BSH is a key mechanistic microbiome target for the development of novel non-antibiotic food additives to improve animal feed production and for the design of new measures to control obesity in humans. However, research on BSH is still in its infancy, particularly in terms of the structural basis of BSH function, which has hampered the development of BSH-based strategies for improving human and animal health. As an initial step towards the structure-function analysis of BSH, C-terminally His-tagged BSH from Lactobacillus salivarius NRRL B-30514 was crystallized in this study. The 1.90 Å resolution crystal structure of L. salivarius BSH was determined by molecular replacement using the structure of Clostridium perfringens BSH as a starting model. It revealed this BSH to be a member of the N-terminal nucleophile hydrolase superfamily. Crystals of apo BSH belonged to space group P21212, with unit-cell parameters a = 90.79, b = 87.35, c = 86.76 Å (PDB entry 5hke). Two BSH molecules packed perfectly as a dimer in one asymmetric unit. Comparative structural analysis of L. salivarius BSH also identified potential residues that contribute to catalysis and substrate specificity.
Organizational Affiliation:
Beijing Key Laboratory for Prevention and Control of Infectious Diseases in Livestock and Poultry, Institute of Animal Husbandry and Veterinary Medicine, Beijing Academy of Agriculture and Forestry Sciences, Beijing 100097, People's Republic of China.