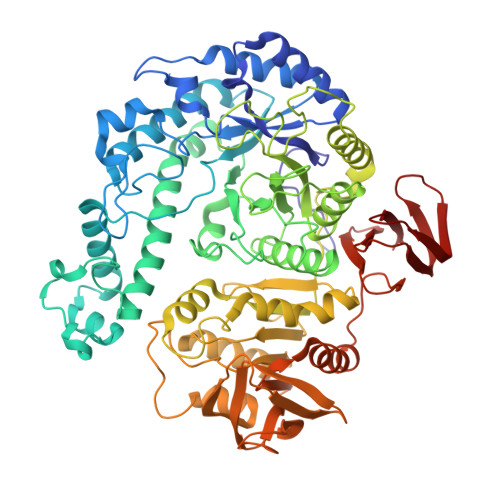
Structure-function relationships in Gan42B, an intracellular GH42 beta-galactosidase from Geobacillus stearothermophilus.
Solomon, H.V., Tabachnikov, O., Lansky, S., Salama, R., Feinberg, H., Shoham, Y., Shoham, G.(2015) Acta Crystallogr D Biol Crystallogr 71: 2433-2448
- PubMed: 26627651
- DOI: https://doi.org/10.1107/S1399004715018672
- Primary Citation of Related Structures:
5DFA - PubMed Abstract:
Geobacillus stearothermophilus T-6 is a Gram-positive thermophilic soil bacterium that contains a battery of degrading enzymes for the utilization of plant cell-wall polysaccharides, including xylan, arabinan and galactan. A 9.4 kb gene cluster has recently been characterized in G. stearothermophilus that encodes a number of galactan-utilization elements. A key enzyme of this degradation system is Gan42B, an intracellular GH42 β-galactosidase capable of hydrolyzing short β-1,4-galactosaccharides into galactose units, making it of high potential for various biotechnological applications. The Gan42B monomer is made up of 686 amino acids, and based on sequence homology it was suggested that Glu323 is the catalytic nucleophile and Glu159 is the catalytic acid/base. In the current study, the detailed three-dimensional structure of wild-type Gan42B (at 2.45 Å resolution) and its catalytic mutant E323A (at 2.50 Å resolution), as determined by X-ray crystallography, are reported. These structures demonstrate that the three-dimensional structure of the Gan42B monomer generally correlates with the overall fold observed for GH42 proteins, consisting of three main domains: an N-terminal TIM-barrel domain, a smaller mixed α/β domain, and the smallest all-β domain at the C-terminus. The two catalytic residues are located in the TIM-barrel domain in a pocket-like active site such that their carboxylic functional groups are about 5.3 Å from each other, consistent with a retaining mechanism. The crystal structure demonstrates that Gan42B is a homotrimer, resembling a flowerpot in general shape, in which each monomer interacts with the other two to form a cone-shaped tunnel cavity in the centre. The cavity is ∼35 Å at the wide opening and ∼5 Å at the small opening and ∼40 Å in length. The active sites are situated at the interfaces between the monomers, so that every two neighbouring monomers participate in the formation of each of the three active sites of the trimer. They are located near the small opening of the cone tunnel, all facing the centre of the cavity. The biological relevance of this trimeric structure is supported by independent results obtained from gel-permeation chromatography. These data and their comparison to the structural data of related GH42 enzymes are used for a more general discussion concerning structure-activity aspects in this GH family.
Organizational Affiliation:
Institute of Chemistry and the Laboratory for Structural Chemistry and Biology, The Hebrew University of Jerusalem, Jerusalem 91904, Israel.