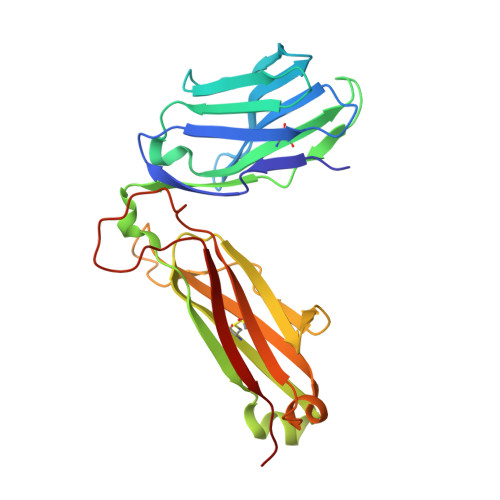
Direct molecular mimicry enables off-target cardiovascular toxicity by an enhanced affinity TCR designed for cancer immunotherapy.
Raman, M.C., Rizkallah, P.J., Simmons, R., Donnellan, Z., Dukes, J., Bossi, G., Le Provost, G.S., Todorov, P., Baston, E., Hickman, E., Mahon, T., Hassan, N., Vuidepot, A., Sami, M., Cole, D.K., Jakobsen, B.K.(2016) Sci Rep 6: 18851-18851
- PubMed: 26758806
- DOI: https://doi.org/10.1038/srep18851
- Primary Citation of Related Structures:
5BRZ, 5BS0 - PubMed Abstract:
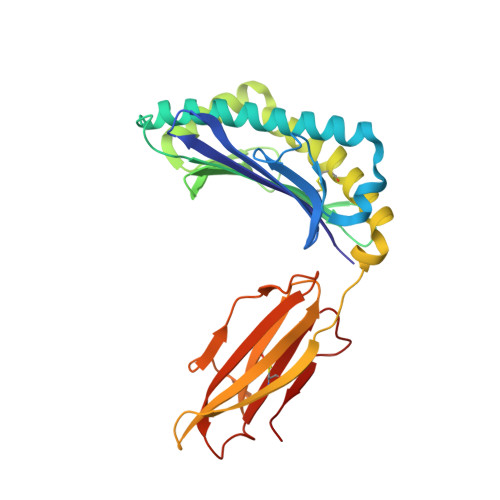
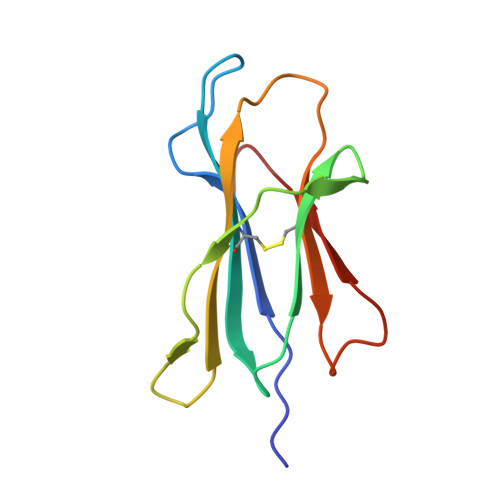
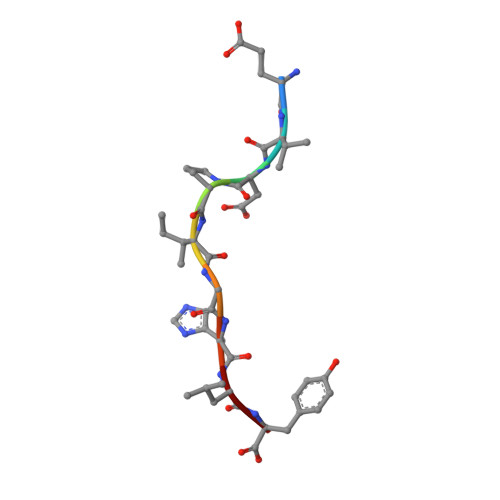
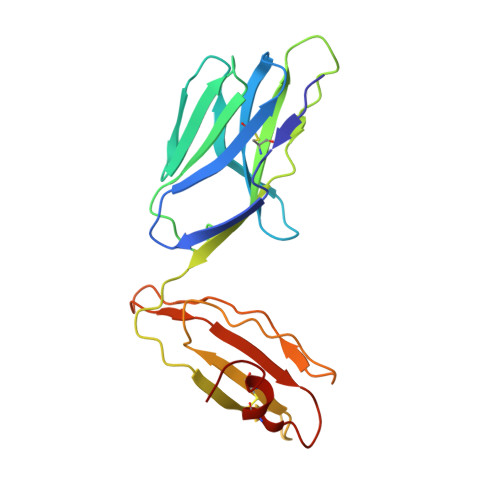
Natural T-cell responses generally lack the potency to eradicate cancer. Enhanced affinity T-cell receptors (TCRs) provide an ideal approach to target cancer cells, with emerging clinical data showing significant promise. Nevertheless, the risk of off target reactivity remains a key concern, as exemplified in a recent clinical report describing fatal cardiac toxicity, following administration of MAGE-A3 specific TCR-engineered T-cells, mediated through cross-reactivity with an unrelated epitope from the Titin protein presented on cardiac tissue. Here, we investigated the structural mechanism enabling TCR cross-recognition of MAGE-A3 and Titin, and applied the resulting data to rationally design mutants with improved antigen discrimination, providing a proof-of-concept strategy for altering the fine specificity of a TCR towards an intended target antigen. This study represents the first example of direct molecular mimicry leading to clinically relevant fatal toxicity, mediated by a modified enhanced affinity TCR designed for cancer immunotherapy. Furthermore, these data demonstrate that self-antigens that are expressed at high levels on healthy tissue should be treated with extreme caution when designing immuno-therapeutics.
Organizational Affiliation:
Immunocore Limited, 101 Park Drive, Milton Park, Abingdon, Oxon, OX14 4RX, United Kingdom.