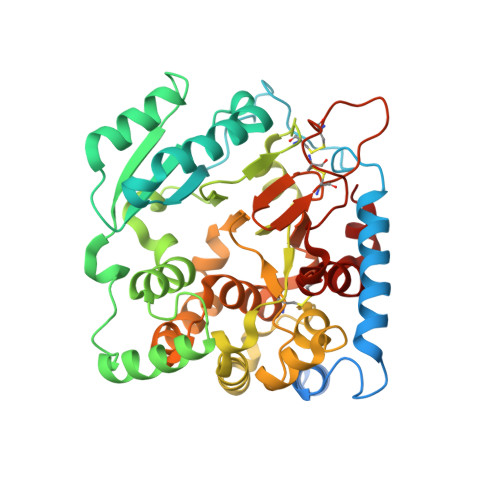
Structure of the Glycosyltransferase Ktr4P from Saccharomyces Cerevisiae
Possner, D.D.D., Claesson, M., Guy, J.E.(2015) PLoS One 10: 36239
- PubMed: 26296208
- DOI: https://doi.org/10.1371/journal.pone.0136239
- Primary Citation of Related Structures:
5A07, 5A08 - PubMed Abstract:
In the yeast Saccharomyces cerevisiae, members of the Kre2/Mnt1 protein family have been shown to be α-1,2-mannosyltransferases or α-1,2-mannosylphosphate transferases, utilising an Mn2+-coordinated GDP-mannose as the sugar donor and a variety of mannose derivatives as acceptors. Enzymes in this family are localised to the Golgi apparatus, and have been shown to be involved in both N- and O-linked glycosylation of newly-synthesised proteins, including cell wall glycoproteins. Our knowledge of the nine proteins in this family is however very incomplete at present. Only one family member, Kre2p/Mnt1p, has been studied by structural methods, and three (Ktr4p, Ktr5p, Ktr7p) are completely uncharacterised and remain classified only as putative glycosyltransferases. Here we use in vitro enzyme activity assays to provide experimental confirmation of the predicted glycosyltransferase activity of Ktr4p. Using GDP-mannose as the donor, we observe activity towards the acceptor methyl-α-mannoside, but little or no activity towards mannose or α-1,2-mannobiose. We also present the structure of the lumenal catalytic domain of S. cerevisiae Ktr4p, determined by X-ray crystallography to a resolution of 2.2 Å, and the complex of the enzyme with GDP to 1.9 Å resolution.
Organizational Affiliation:
Department of Medical Biochemistry and Biophysics, Karolinska Institutet, Tomtebodavägen 6, S 171-77, Stockholm, Sweden.