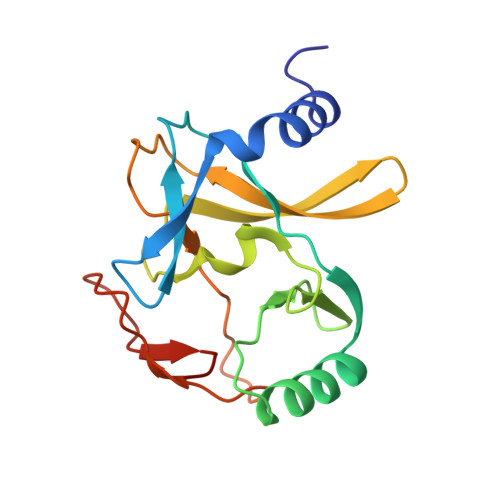
Evolving Catalytic Properties of the MLL Family SET Domain.
Zhang, Y., Mittal, A., Reid, J., Reich, S., Gamblin, S.J., Wilson, J.R.(2015) Structure 23: 1921-1933
- PubMed: 26320581
- DOI: https://doi.org/10.1016/j.str.2015.07.018
- Primary Citation of Related Structures:
4Z4P - PubMed Abstract:
Methylation of histone H3 lysine-4 is a hallmark of chromatin associated with active gene expression. The activity of H3K4-specific modification enzymes, in higher eukaryotes the MLL (or KMT2) family, is tightly regulated. The MLL family has six members, each with a specialized function. All contain a catalytic SET domain that associates with a core multiprotein complex for activation. These SET domains segregate into three classes that correlate with the arrangement of targeting domains that populate the rest of the protein. Here we show that, unlike MLL1, the MLL4 SET domain retains significant activity without the core complex. We also present the crystal structure of an inactive MLL4-tagged SET domain construct and describe conformational changes that account for MLL4 intrinsic activity. Finally, our structure explains how the MLL SET domains are able to add multiple methyl groups to the target lysine, despite having the sequence characteristics of a classical monomethylase.
Organizational Affiliation:
The Francis Crick Institute, Mill Hill Laboratory, London NW7 1AA, UK.