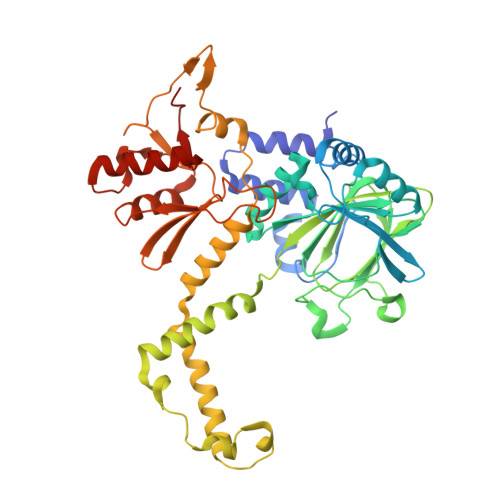
Structure of the JmjC domain-containing protein NO66 complexed with ribosomal protein Rpl8.
Wang, C., Zhang, Q., Hang, T., Tao, Y., Ma, X., Wu, M., Zhang, X., Zang, J.(2015) Acta Crystallogr D Biol Crystallogr 71: 1955-1964
- PubMed: 26327385
- DOI: https://doi.org/10.1107/S1399004715012948
- Primary Citation of Related Structures:
4Y33, 4Y3O, 4Y4R - PubMed Abstract:
The JmjC domain-containing proteins belong to a large family of oxygenases possessing distinct substrate specificities which are involved in the regulation of different biological processes, such as gene transcription, RNA processing and translation. Nucleolar protein 66 (NO66) is a JmjC domain-containing protein which has been reported to be a histone demethylase and a ribosome protein 8 (Rpl8) hydroxylase. The present biochemical study confirmed the hydroxylase activity of NO66 and showed that oligomerization is required for NO66 to efficiently catalyze the hydroxylation of Rpl8. The structures of NO66(176-C) complexed with Rpl8(204-224) in a tetrameric form and of the mutant protein M2 in a dimeric form were solved. Based on the results of structural and biochemical analyses, the consensus sequence motif NHXH recognized by NO66 was confirmed. Several potential substrates of NO66 were found by a BLAST search according to the consensus sequence motif. When binding to substrate, the relative positions of each subunit in the NO66 tetramer shift. Oligomerization may facilitate the motion of each subunit in the NO66 tetramer and affect the catalytic activity.
Organizational Affiliation:
Hefei National Laboratory for Physical Sciences at Microscale and School of Life Sciences, Collaborative Innovation Center of Chemistry for Life Science, University of Science and Technology of China, 96 Jinzhai Road, Hefei, Anhui 230026, People's Republic of China.