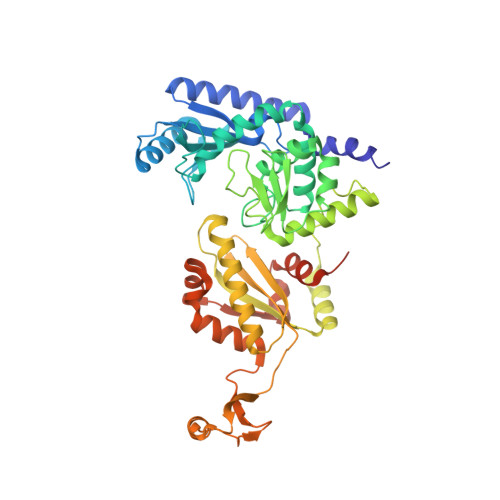
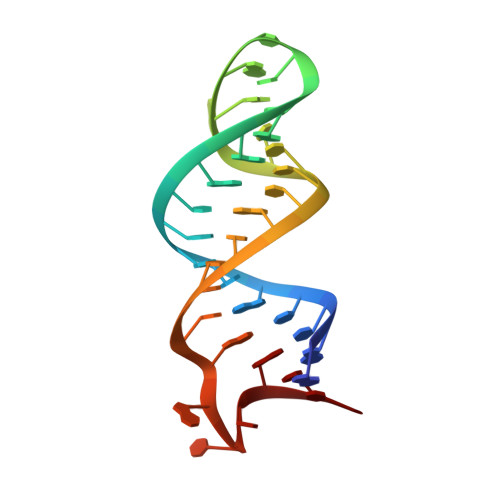
On-Enzyme Refolding Permits Small RNA and tRNA Surveillance by the CCA-Adding Enzyme.
Kuhn, C.D., Wilusz, J.E., Zheng, Y., Beal, P.A., Joshua-Tor, L.(2015) Cell 160: 644-658
- PubMed: 25640237
- DOI: https://doi.org/10.1016/j.cell.2015.01.005
- Primary Citation of Related Structures:
4X4N, 4X4O, 4X4P, 4X4Q, 4X4R, 4X4S, 4X4T, 4X4U, 4X4V, 4X4W - PubMed Abstract:
Transcription in eukaryotes produces a number of long noncoding RNAs (lncRNAs). Two of these, MALAT1 and Menβ, generate a tRNA-like small RNA in addition to the mature lncRNA. The stability of these tRNA-like small RNAs and bona fide tRNAs is monitored by the CCA-adding enzyme. Whereas CCA is added to stable tRNAs and tRNA-like transcripts, a second CCA repeat is added to certain unstable transcripts to initiate their degradation. Here, we characterize how these two scenarios are distinguished. Following the first CCA addition cycle, nucleotide binding to the active site triggers a clockwise screw motion, producing torque on the RNA. This ejects stable RNAs, whereas unstable RNAs are refolded while bound to the enzyme and subjected to a second CCA catalytic cycle. Intriguingly, with the CCA-adding enzyme acting as a molecular vise, the RNAs proofread themselves through differential responses to its interrogation between stable and unstable substrates.
Organizational Affiliation:
W.M. Keck Structural Biology Laboratory, Howard Hughes Medical Institute, Cold Spring Harbor Laboratory, 1 Bungtown Road, Cold Spring Harbor, NY 11724, USA.