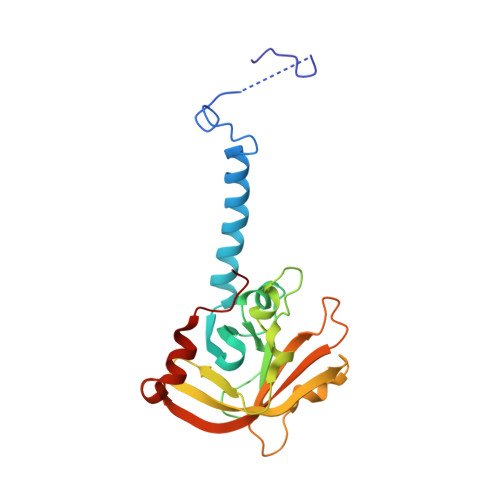
Molecular basis of Streptococcus mutans sortase A inhibition by the flavonoid natural product trans-chalcone.
Wallock-Richards, D.J., Marles-Wright, J., Clarke, D.J., Maitra, A., Dodds, M., Hanley, B., Campopiano, D.J.(2015) Chem Commun (Camb) 51: 10483-10485
- PubMed: 26029850
- DOI: https://doi.org/10.1039/c5cc01816a
- Primary Citation of Related Structures:
4TQX - PubMed Abstract:
Sortase A (SrtA) from Gram positive pathogens is an attractive target for inhibitors due to its role in the attachment of surface proteins to the cell wall. We found that the plant natural product trans-chalcone inhibits Streptococcus mutans SrtA in vitro and also inhibited S. mutans biofilm formation. Mass spectrometry revealed that the trans-chalcone forms a Michael addition adduct with the active site cysteine. The X-ray crystal structure of the SrtA H139A mutant provided new insights into substrate recognition by the sortase family. Our study suggests that chalcone flavonoids have potential as sortase-specific oral biofilm inhibitors.
Organizational Affiliation:
EastChem School of Chemistry, The University of Edinburgh, David Brewster Road, Edinburgh, EH9 3FJ, UK. Dominic.Campopiano@ed.ac.uk.