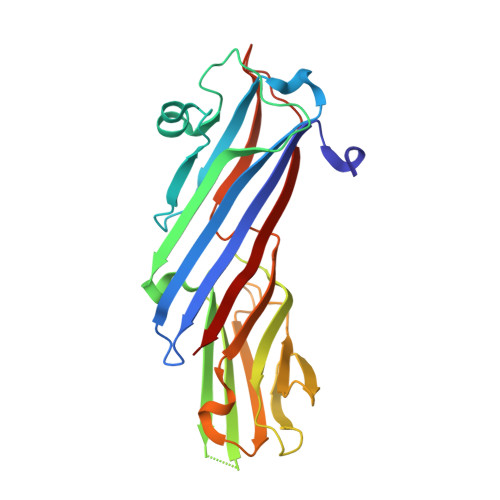
Structure of the bovine COPI delta subunit mu homology domain at 2.15 angstrom resolution.
Lahav, A., Rozenberg, H., Parnis, A., Cassel, D., Adir, N.(2015) Acta Crystallogr D Biol Crystallogr 71: 1328-1334
- PubMed: 26057672
- DOI: https://doi.org/10.1107/S1399004715006203
- Primary Citation of Related Structures:
4O8Q - PubMed Abstract:
The heptameric COPI coat (coatomer) plays an essential role in vesicular transport in the early secretory system of eukaryotic cells. While the structures of some of the subunits have been determined, that of the δ-COP subunit has not been reported to date. The δ-COP subunit is part of a subcomplex with structural similarity to tetrameric clathrin adaptors (APs), where δ-COP is the structural homologue of the AP μ subunit. Here, the crystal structure of the μ homology domain (MHD) of δ-COP (δ-MHD) obtained by phasing using a combined SAD-MR method is presented at 2.15 Å resolution. The crystallographic asymmetric unit contains two monomers that exhibit short sections of disorder, which may allude to flexible regions of the protein. The δ-MHD is composed of two subdomains connected by unstructured linkers. Comparison between this structure and those of known MHD domains from the APs shows significant differences in the positions of specific loops and β-sheets, as well as a more general change in the relative positions of the protein subdomains. The identified difference may be the major source of cargo-binding specificity. Finally, the crystal structure is used to analyze the potential effect of the I422T mutation in δ-COP previously reported to cause a neurodegenerative phenotype in mice.
Organizational Affiliation:
Schulich Faculty of Chemistry, Technion, Haifa 32000, Israel.