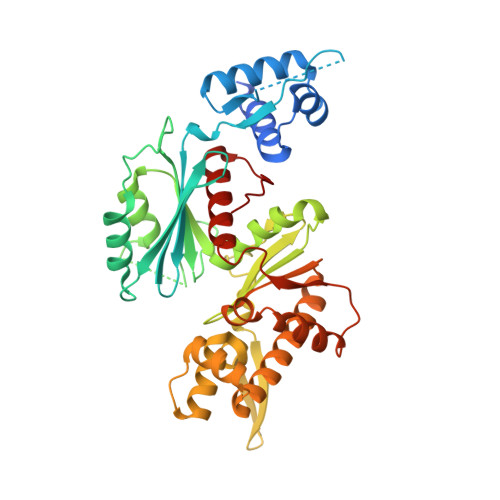
Structure-Function Studies of the Staphylococcal Methicillin Resistance Antirepressor MecR2.
Arede, P., Botelho, T., Guevara, T., Uson, I., Oliveira, D.C., Gomis-Ruth, F.X.(2013) J Biol Chem 288: 21267-21278
- PubMed: 23733184
- DOI: https://doi.org/10.1074/jbc.M112.448134
- Primary Citation of Related Structures:
4IJA - PubMed Abstract:
Methicillin resistance in Staphylococcus aureus is elicited by the MecI-MecR1-MecA axis encoded by the mec locus. Recently, MecR2 was also identified as a regulator of mec through binding of the methicillin repressor, MecI. Here we show that plasmid-encoded full-length MecR2 restores resistance in a sensitive S. aureus mecR2 deletion mutant of the resistant strain N315. The crystal structure of MecR2 reveals an N-terminal DNA-binding domain, an intermediate scaffold domain, and a C-terminal dimerization domain that contributes to oligomerization. The protein shows structural similarity to ROK (repressors, open reading frames, and kinases) family proteins, which bind DNA and/or sugar molecules. We found that functional cell-based assays of three point mutants affecting residues participating in sugar binding in ROK proteins had no effect on the resistance phenotype. By contrast, MecR2 bound short double-stranded DNA oligonucleotides nonspecifically, and a deletion mutant affecting the N-terminal DNA-binding domain showed a certain effect on activity, thus contributing to resistance less than the wild-type protein. Similarly, a deletion mutant, in which a flexible segment of intermediate scaffold domain had been replaced by four glycines, significantly reduced MecR2 function, thus indicating that this domain may likewise be required for activity. Taken together, these results provide the structural basis for the activity of a methicillin antirepressor, MecR2, which would sequester MecI away from its cognate promoter region and facilitate its degradation.
Organizational Affiliation:
the Center for Microbiological Resources, Department of Life Sciences, Faculdade de Ciências e Tecnologia, Universidade Nova de Lisboa, Quinta da Torre, P-2829-516 Caparica, Portugal, and.